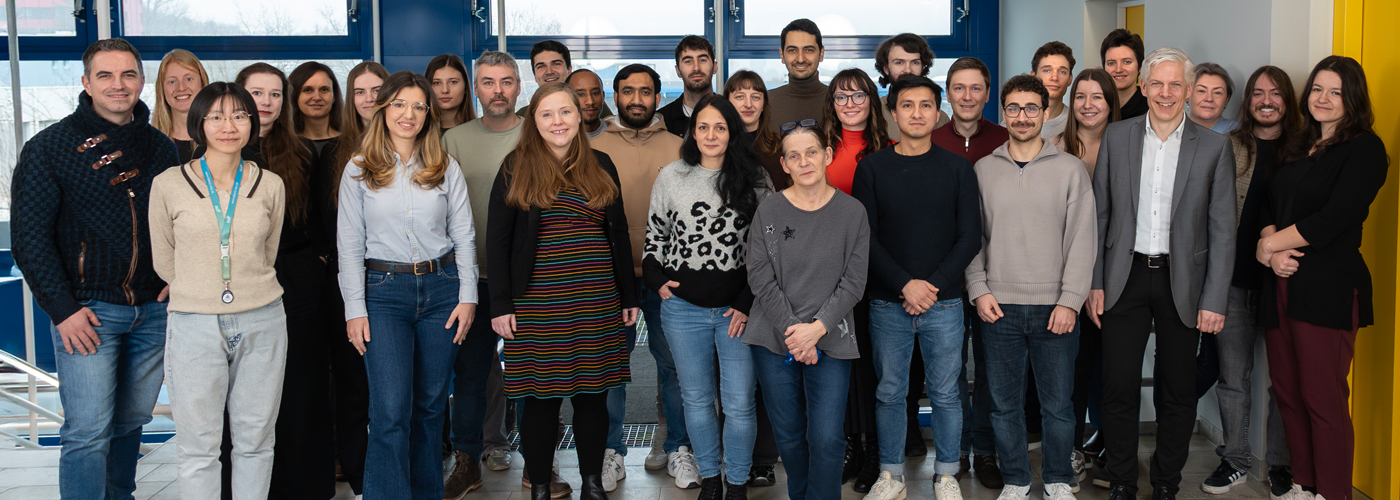
Unsere Inspiration ist die Anpassungsfähigkeit von Organismen und den Materialien, aus denen sie aufgebaut sind, an wechselnde Umweltbedingungen. Pflanzen passen ihr Wachstum an die Lichtverhältnisse an, Bakterien entwickeln Resistenzen gegen Antibiotika oder Knochen werden durch Belastung stärker. Grundlage für diese Anpassungsfähigkeit ist eine faszinierende Signalverarbeitung der Organismen: Durch molekulare Sensoren werden Umweltbedingungen wahrgenommen, die Signale werden prozessiert und mit dem genetischen Programm des Organismus integriert, um am Ende eine passgenaue Reaktion auszulösen.
In unserer Forschung verwenden wir diese molekularen informationsverarbeitenden Mechanismen, um die Funktion und Eigenschaften von Zellen und Materialien gezielt zu steuern. Dies eröffnet neuartige Möglichkeiten in der grundladen- und anwendungsorientierten Forschung.
Mehr Informationen finden Sie auf unserer englischsprachigen Seite.

Mitarbeiter/innen
Publikationen
Kolb, L. | Allazetta, S. | Karlsson, M. | Girgin, M. | Weber, Wilfried | Lutolf, M. P.
DOI:
As the field of tissue engineering develops, methods for screening combinations of signals for their effects on stem cell behavior are needed. We introduce a microgel-based screening platform for testing combinations of in situ-generated proteins on stem cell fate in ultrahigh-throughput. Compartmentalizing individual sets of growth factors was addressed by encapsulating aggregates of stable recombinant cell lines secreting individual glycoproteins into microgels through an on-chip polymerization. When these 'microniches' are cultured with a cell type of interest, fluorescence reporters indicate positive niches that perform the desired function, and the underlying producer cell lines of these selected microniches are analyzed by barcoded RNA sequencing. The microniche-based screening work-flow was validated via a model system based on engineered mammalian cells expressing yellow fluorescent protein (YFP) upon anti-inflammatory cytokine interleukin 4 (IL4)-based activation. © 2019 The Royal Society of Chemistry.
Hörner, M. | Raute, K. | Hummel, B. | Madl, J. | Creusen, G. | Thomas, O. S. | Christen, E. H. | Hotz, N. | Gübeli, R. J. | Engesser, R. | Rebmann, B. | Lauer, J. | Rolauffs, B. | Timmer, J. | Schamel, W. W. A. | Pruszak, J. | Römer, W. | Zurbriggen, M. D. | Friedrich, C. | Walther, A. | Minguet, S. | Sawarkar, R. | Weber, Wilfried
DOI:
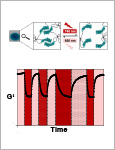
Interrogation and control of cellular fate and function using optogenetics is providing revolutionary insights into biology. Optogenetic control of cells is achieved by coupling genetically encoded photoreceptors to cellular effectors and enables unprecedented spatiotemporal control of signaling processes. Here, a fast and reversibly switchable photoreceptor is used to tune the mechanical properties of polymer materials in a fully reversible, wavelength-specific, and dose- and space-controlled manner. By integrating engineered cyanobacterial phytochrome 1 into a poly(ethylene glycol) matrix, hydrogel materials responsive to light in the cell-compatible red/far-red spectrum are synthesized. These materials are applied to study in human mesenchymal stem cells how different mechanosignaling pathways respond to changing mechanical environments and to control the migration of primary immune cells in 3D. This optogenetics-inspired matrix allows fundamental questions of how cells react to dynamic mechanical environments to be addressed. Further, remote control of such matrices can create new opportunities for tissue engineering or provide a basis for optically stimulated drug depots. © 2019 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim
Mühlhäuser, W. W. D. | Weber, Wilfried | Radziwill, G.
DOI:
Subcellular localization of signal molecules is a hallmark in organizing the signaling network. OpEn-Tag is a modular optogenetic endomembrane targeting toolbox that allows alteration of the localization and therefore the activity of signaling processes with the spatiotemporal resolution of optogenetics. OpEn-Tag is a two-component system employing (1) a variety of targeting peptides fused to and thereby dictating the localization of mCherry-labeled cryptochrome 2 binding protein CIBN toward distinct endomembranes and (2) the cytosolic, fluorescence-labeled blue light photoreceptor cryptochrome 2 as a customizable building block that can be fused to proteins of interest. The combination of OpEn-Tag with growth factor stimulation or the use of two membrane anchor sequences allows investigation of multilayered signal transduction processes as demonstrated here for the protein kinase AKT. © 2019 American Chemical Society.
Thomas, O. S. | Weber, Wilfried
DOI:
The exploitation of nanosized materials for the delivery of therapeutic agents is already a clinical reality and still holds unrealized potential for the treatment of a variety of diseases. This review discusses physiological barriers a nanocarrier must overcome in order to reach its target, with an emphasis on cancer nanomedicine. Stages of delivery include residence in the blood stream, passive accumulation by virtue of the enhanced permeability and retention effect, diffusion within the tumor lesion, cellular uptake, and arrival at the site of action. We also briefly outline strategies for engineering nanoparticles to more efficiently overcome these challenges: Increasing circulation half-life by shielding with hydrophilic polymers, such as PEG, the limitations of PEG and potential alternatives, targeting and controlled activation approaches. Future developments in these areas will allow us to harness the full potential of nanomedicine. © Copyright © 2019 Thomas and Weber.
Wagner, H. J. | Engesser, R. | Ermes, K. | Geraths, C. | Timmer, J. | Weber, Wilfried
DOI:
Synthetic biology applies engineering concepts to build cells that perceive and process information. Examples include cells engineered to perform basic digital or analog computation. These circuits serve as basis for the construction of complex integrated cellular networks that offer manifold applications in fundamental and applied research. Here, we introduce the concept of using design approaches and molecular tools applied in synthetic biology for the construction of interconnected biohybrid materials systems with information processing functionality. We validate this concept by modularly assembling protein and polymer building blocks to generate stimulus-responsive materials. Guided by a quantitative mathematical model, we next interconnect these materials into a materials system that acts as both a signal detector and as an amplifier based on a built-in positive feedback loop. The functionality and versatility of this materials system is demonstrated by the detection of enzymatic activities and drugs. The modular design concept presented here thus represents a blueprint for integrating synthetic biology-inspired information-processing circuits into polymer materials. As integrated sensors and actuators, the resulting smart materials systems could provide novel solutions with broad perspectives in research and development. © 2018 Elsevier Ltd
Wagner, H. J. | Kemmer, S. | Engesser, R. | Timmer, J. | Weber, Wilfried
DOI:
Feedforward and feedback loops are key regulatory elements in cellular signaling and information processing. Synthetic biology exploits these elements for the design of molecular circuits that enable the reprogramming and control of specific cellular functions. These circuits serve as a basis for the engineering of complex cellular networks, opening the door for numerous medical and biotechnological applications. Here, a similar principle is applied. Feedforward and positive feedback circuits are incorporated into biohybrid polymer materials in order to develop signal-sensing and signal-processing devices. This concept is exemplified by the detection of the proteolytic activity of the botulinum neurotoxin A. To this aim, site-specific proteases are incorporated into receiver, transmitter, and output materials, and their release, diffusion, and/or activation are wired according to a feedforward or a positive feedback circuit. The development of a quantitative mathematical model enables analysis and comparison of the performance of both systems. The flexible design could be easily adapted to detect other toxins or molecules of interest. Furthermore, cellular signaling or gene regulatory pathways could provide additional blueprints for the development of novel biohybrid circuits. Such information-processing, material-embedded biological circuits hold great promise for a variety of analytical, medical, or biotechnological applications. © 2018 The Authors. Published by WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim
Wagner, H. J. | Weber, Wilfried
DOI:
The engineering of enzymes for the purpose of controlling their activity represents a valuable approach to address challenges in both fundamental and applied research. Here, we describe and compare different design strategies for the generation of a human rhinovirus-14 (HRV14) 3C protease-inducible caspase-3 (CASP3). We exemplify the application potential of the resulting protease by controlling the activity of a synthetic enzyme cascade, which represents an important motif for the design of artificial signal transduction networks. In addition, we use our engineered CASP3 to characterize the effect of aspartate mutations on enzymatic activity. Besides the identification of mutations that render the enzyme inactive, we find the CASP3-D192E mutant (aspartate-to-glutamate exchange at position 192) to be inaccessible for 3C protease-mediated cleavage. This indicates a structural change of CASP3 that goes beyond a slight misalignment of the catalytic triad. This study could inspire the design of additional engineered proteases that could be used to unravel fundamental research questions or to expand the collection of biological parts for the design of synthetic signaling pathways. © 2019 by the authors.
Yousefi, O. S. | Günther, M. | Hörner, M. | Chalupsky, J. | Wess, M. | Brandl, S. M. | Smith, R. W. | Fleck, C. | Kunkel, T. | Zurbriggen, M. D. | Höfer, T. | Weber, Wilfried | Schamel, W. W.
DOI:
The immune system distinguishes between self and foreign antigens. The kinetic proofreading (KPR) model proposes that T cells discriminate self from foreign ligands by the different ligand binding half-lives to the T cell receptor (TCR). It is challenging to test KPR as the available experimental systems fall short of only altering the binding half-lives and keeping other parameters of the interaction unchanged. We engineered an optogenetic system using the plant photoreceptor phytochrome B (PhyB) as a ligand to selectively control the dynamics of ligand binding to the TCR by light. This opto-ligand-TCR system was combined with the unique property of PhyB to continuously cycle between the binding and non-binding states under red light, with the light intensity determining the cycling rate and thus the binding duration. Mathematical modeling of our experimental datasets showed that indeed the ligand-TCR interaction half-life is the decisive factor for activating downstream TCR signaling, substantiating KPR. © Yousefi et al.
Baaske, J. | Gonschorek, P. | Engesser, R. | Dominguez-Monedero, A. | Raute, K. | Fischbach, P. | Müller, K. | Cachat, E. | Schamel, W. W. A. | Minguet, S. | Davies, J. A. | Timmer, J. | Weber, Wilfried | Zurbriggen, M. D.
DOI:
Optogenetic switches are emerging molecular tools for studying cellular processes as they offer higher spatiotemporal and quantitative precision than classical, chemical-based switches. Light-controllable gene expression systems designed to upregulate protein expression levels meanwhile show performances superior to their chemical-based counterparts. However, systems to reduce protein levels with similar efficiency are lagging behind. Here, we present a novel two-component, blue light-responsive optogenetic OFF switch (‘Blue-OFF’), which enables a rapid and quantitative down-regulation of a protein upon illumination. Blue-OFF combines the first light responsive repressor KRAB-EL222 with the protein degradation module B-LID (blue light-inducible degradation domain) to simultaneously control gene expression and protein stability with a single wavelength. Blue-OFF thus outperforms current optogenetic systems for controlling protein levels. The system is described by a mathematical model which aids in the choice of experimental conditions such as light intensity and illumination regime to obtain the desired outcome. This approach represents an advancement of dual-controlled optogenetic systems in which multiple photosensory modules operate synergistically. As exemplified here for the control of apoptosis in mammalian cell culture, the approach opens up novel perspectives in fundamental research and applications such as tissue engineering. © 2018, The Author(s).
Beyer, H. M. | Engesser, R. | Hörner, M. | Koschmieder, J. | Beyer, P. | Timmer, J. | Zurbriggen, M. D. | Weber, Wilfried
DOI:
Synthetic biology applies engineering concepts to build cellular systems that perceive and process information. This is achieved by assembling genetic modules according to engineering design principles. Recent advance in the field has contributed optogenetic switches for controlling diverse biological functions in response to light. Here, the concept is introduced to apply synthetic biology switches and design principles for the synthesis of multi-input-processing materials. This is exemplified by the synthesis of a materials system that counts light pulses. Guided by a quantitative mathematical model, functional synthetic biology-derived modules are combined into a polymer framework resulting in a biohybrid materials system that releases distinct output molecules specific to the number of input light pulses detected. Further demonstration of modular extension yields a light pulse-counting materials system to sequentially release different enzymes catalyzing a multistep biochemical reaction. The resulting smart materials systems can provide novel solutions as integrated sensors and actuators with broad perspectives in fundamental and applied research. © 2018 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim

