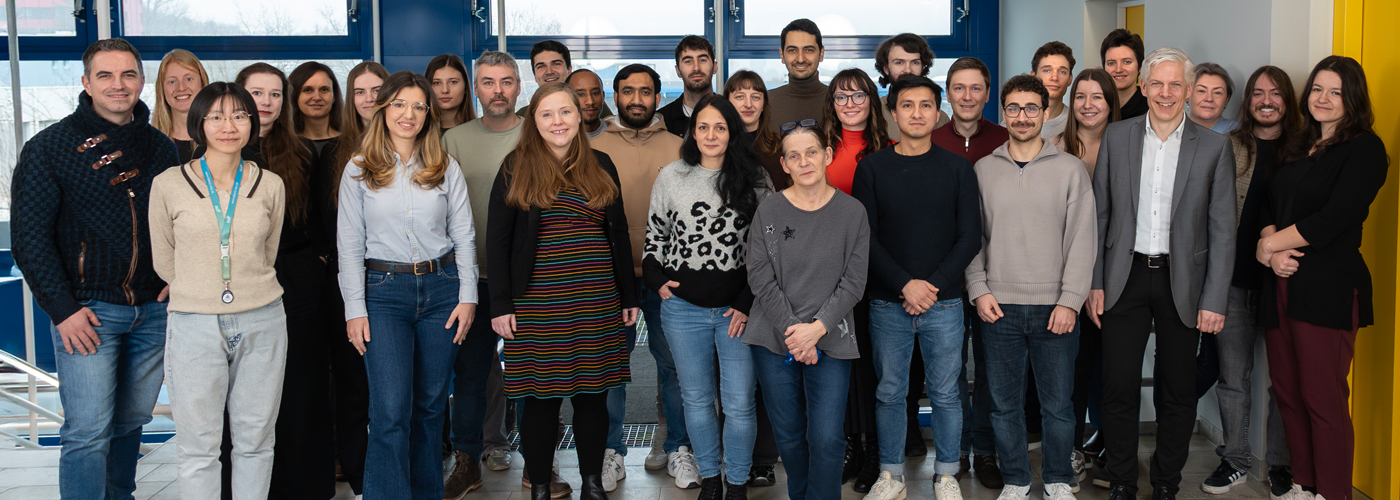
We engineer cells and materials that communicate and process information through synthetic biology
Our inspiration is the ability of organisms and the materials they are made of to adapt to dynamic environmental conditions. Plants adapt growth to light conditions; bacteria develop resistance against antibiotics or bones get stronger when exercised. The basis for this ability to adapt is a fascinating information processing machinery of the organisms: Environmental conditions are captured by molecular sensors, then the signals are processed and integrated with genetic programs to finally yield a targeted response.
In our research, we engineer nature’s molecular sensing, processing, and actuation machinery in order to precisely control the function and properties of cells and materials. We apply these newly developed technologies in different fields of fundamental and applied research.

Team Members











Research
Stimulus-responsive and Information-processing (living) Materials

We develop and apply stimulus-responsive and information-processing biohybrid polymer materials. To this aim, we functionally couple synthetic biological molecular sensors and switches to polymer materials. By wiring these switches according to topologies inspired by electronic circuits, we engineer materials that perform fundamental computational operations. Examples of our work include:
- We engineered a hydrogel based on a bacteria-derived photoreceptor which allows the light-responsive, fully reversibly tuning of its mechanical properties. We applied this hydrogel as extracellular matrix to analyze the impact of dynamic mechanical environments on transcriptome-wide responses in mesenchymal stem cells or on the migration of T-lymphocytes.
See Hörner et al. Advanced Materials 2019 - We integrated synthetic biological switches with polymer materials into a circuit inspired by an electronic counter. The resulting material system was able to count the number of input light pulses and to release different output as a function of the number of light pulses detected. We applied this system to sequentially release different biocatalysts to drive a two-step biochemical reaction.
See Beyer et al., Advanced Materials 2018 - We developed PenTag, a protein tag for the spontaneous, covalent coupling of proteins to ampicillin-functionalized molecules such as dyes, polymers, or solid supports. Based on this strategy, we engineered and assembled material modules to function as encoder for processing different combinations of biochemical input stimuli.
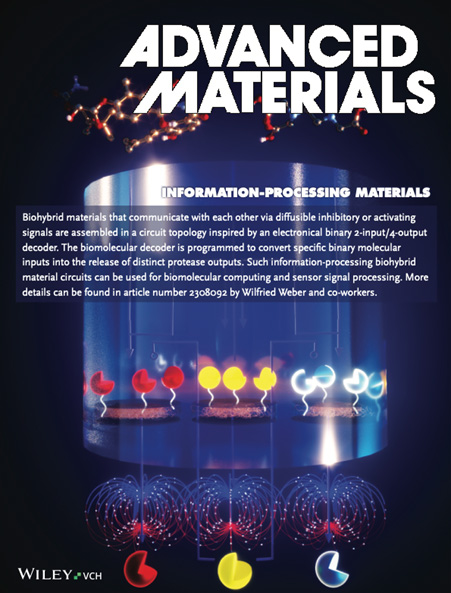
See Mohsenin et al., Advanced Functional Materials 2024 - By engineering modular protease-based switches that can either be activated or repressed, we develop information-processing biohybrid circuits that process binary biomolecular information according to a circuit inspired by electronic decoders. Such circuits can be applied to process and interpret biochemical sensor information for advanced diagnostic applications.
See Mohsenin et al., Advanced Materials 2024
Molecular optogenetics to control cell fate and function
We develop and apply molecular optogenetic tools to control cell fate and function with unprecedented spatial and temporal precision in a dose-dependent and highly specific manner. To this aim, we engineer plant- and bacteria-derived photoreceptors and functionally couple them to proteins involved in cell signaling and gene expression. Examples of our work include:
- Light-inducible formation of liquid or gel-like transcription factor condensates in mammalian cells and mice. We demonstrate that liquid “transcription factor droplets” show a several-fold higher activity in inducing transgene expression compared to native transcription factors. Further, gel-like transcription factor condensates were shown to correlate with decreased transcriptional activation thus providing a materials-based layer of controlling gene expression.
See Schneider et al., Science Advances 2021 and Fischer et al., Small 2024 - Light-guided adeno-associated viral (AAV) vectors. We engineered a light-responsive tropism into AAVs which allows us to selectively transfer genetic information into single cells or to transduce different cells within one culture with different transgenes.
See Hörner et al., Science Advances 2021
Our group is running www.optobase.org, the most comprehensive database on molecular optogenetics. Have a look and discover the amazing opportunities in controlling biology with light!
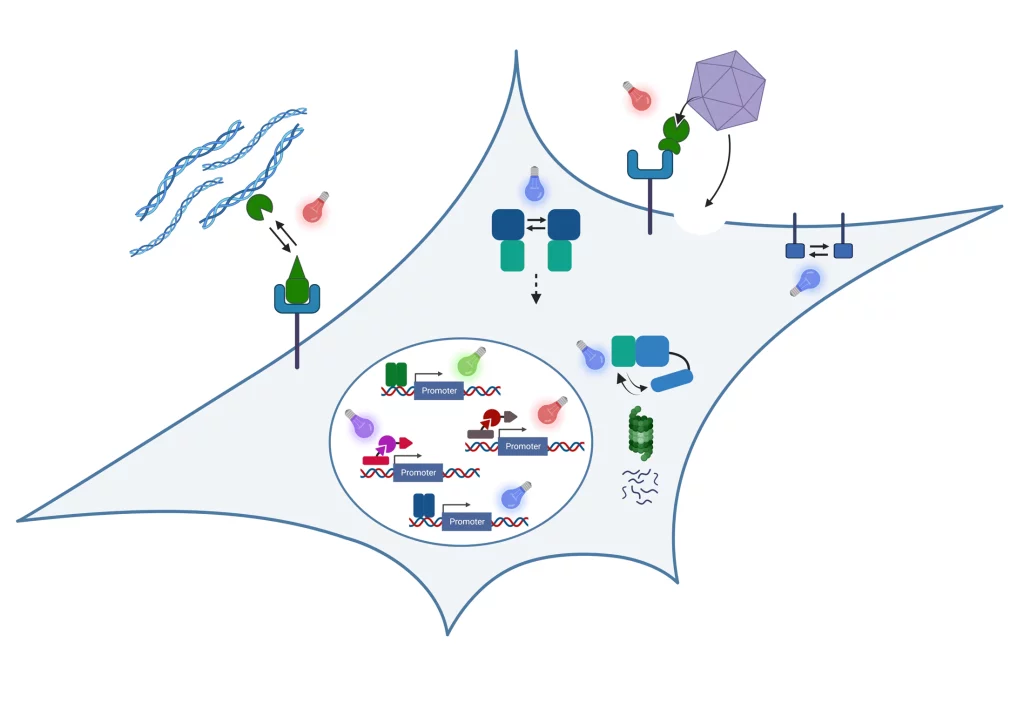
Biosensors
We integrate natural and engineered molecular sensors for drugs, metabolites or nucleic acids into suitable readout formats for the fast and sensitive quantification of such substances. Together with collaboration partners, we develop biosensor systems for different application fields:
Open Positions
We are always excited to meet curious and creative scientists passionate about synthetic biology, optogenetics, and engineered living materials. If you would like to shape the future of biobased and living materials with us, we warmly welcome your spontaneous application for a PhD thesis or Postdoc position!
Projects and Partners
We perform collaborative research in materials-oriented synthetic biology within interdisciplinary research consortia
STEADY
Within the ERC Advanced Grant STEADY, we develop concepts for dynamically controlling the properties of engineered living materials by advanced synthetic genetic circuits.
LoopOfFun
We coordinate the European Innovation Council (EIC)-funded consortium LoopOfFun in which we aim at developing a platform for the rapid development of industry-scale, one-step, simple casting-based manufacturing processes for fungal mycelia composites. We jointly work towards this goal with our consortium partners:
- Prof. Roman Jerala, National Institute of Chemistry, Ljubljana, Slovenia
- Dr. Achim Weber, Fraunhofer IGB, Stuttgart, Germany
- Prof. Arnold Driessen, University of Groningen, The Netherlands
- Carlotta Borgato and Jan Boelen, Atelier LUMA, Arles, France
DELIVER
In the project DELIVER funded by the Carl-Zeiss-Foundation, we collaborate towards the data-driven engineering of sustainable living materials. We combine synthetic biology with materials sciences and data-driven approaches to design bio-based composite materials with custom-tailored structural properties for construction applications. Within deliver, we collaborate with the following partners:
- Prof. Thomas Speck, University of Freiburg, Germany
- Dr. Clemens Kreutz, University Hospital Freiburg, Germany
BILLARD
We coordinate the BILLARD project funded by the Federal Ministry of Education and Research (BMBF) within the funding line “Biologization of Technology”, we collaborate with PD Dr. Felicitas Bucher from the Clinic of Ophtamology at the University Hospital Freiburg on the development of novel intraocular drug delivery devices.
CIBSS – Centre for Integrative Biological Signalling Studies
We are member of the Cluster of Excellence CIBSS in which we perform research on novel optogenetic technologies to control signaling reactions in mammalian cells. We mainly collaborate with Prof. Dr. Jens Timmer on the model-based design of synthetic biological switches and networks and with Prof. Dr. Wolfgang Schamel on controlling immunological processes such as T cell activation via optogenetics.
Publications

Schneider, N. | Wieland, F. G. | Kong, D. | Fischer, A. A. M. | Hörner, M. | Timmer, J. | Ye, H. | Weber, Wilfried
DOI:
Light-inducible gene switches represent a key strategy for the precise manipulation of cellular events in fundamental and applied research. However, the performance of widely used gene switches is limited due to low tissue penetrance and possible phototoxicity of the light stimulus. To overcome these limitations, we engineer optogenetic synthetic transcription factors to undergo liquid-liquid phase separation in close spatial proximity to promoters. Phase separation of constitutive and optogenetic synthetic transcription factors was achieved by incorporation of intrinsically disordered regions. Supported by a quantitative mathematical model, we demonstrate that engineered transcription factor droplets form at target promoters and increase gene expression up to fivefold. This increase in performance was observed in multiple mammalian cells lines as well as in mice following in situ transfection. The results of this work suggest that the introduction of intrinsically disordered domains is a simple yet effective means to boost synthetic transcription factor activity. Copyright © 2021 The Authors, some rights reserved.
Hörner, M. | Jerez-Longres, C. | Hudek, A. | Hook, S. | Yousefi, O. S. | Schamel, W. W. A. | Hörner, C. | Zurbriggen, M. D. | Ye, H. | Wagner, H. J. | Weber, Wilfried
DOI:
Methodologies for the controlled delivery of genetic information into target cells are of utmost importance for genetic engineering in both fundamental and applied research. However, available methods for efficient gene transfer into user-selected or even single cells suffer from low throughput, the need for complicated equipment, high invasiveness, or side effects by off-target viral uptake. Here, we engineer an adeno-associated viral (AAV) vector system that transfers genetic information into native target cells upon illumination with cell-compatible red light. This OptoAAV system allows adjustable and spatially resolved gene transfer down to single-cell resolution and is compatible with different cell lines and primary cells. Moreover, the sequential application of multiple OptoAAVs enables spatially resolved transduction with different transgenes. The approach presented is likely extendable to other classes of viral vectors and is expected to foster advances in basic and applied genetic research. Copyright © 2021 The Authors, some rights reserved.
Hörner, M. | Raute, K. | Hummel, B. | Madl, J. | Creusen, G. | Thomas, O. S. | Christen, E. H. | Hotz, N. | Gübeli, R. J. | Engesser, R. | Rebmann, B. | Lauer, J. | Rolauffs, B. | Timmer, J. | Schamel, W. W. A. | Pruszak, J. | Römer, W. | Zurbriggen, M. D. | Friedrich, C. | Walther, A. | Minguet, S. | Sawarkar, R. | Weber, Wilfried
DOI:
Interrogation and control of cellular fate and function using optogenetics is providing revolutionary insights into biology. Optogenetic control of cells is achieved by coupling genetically encoded photoreceptors to cellular effectors and enables unprecedented spatiotemporal control of signaling processes. Here, a fast and reversibly switchable photoreceptor is used to tune the mechanical properties of polymer materials in a fully reversible, wavelength-specific, and dose- and space-controlled manner. By integrating engineered cyanobacterial phytochrome 1 into a poly(ethylene glycol) matrix, hydrogel materials responsive to light in the cell-compatible red/far-red spectrum are synthesized. These materials are applied to study in human mesenchymal stem cells how different mechanosignaling pathways respond to changing mechanical environments and to control the migration of primary immune cells in 3D. This optogenetics-inspired matrix allows fundamental questions of how cells react to dynamic mechanical environments to be addressed. Further, remote control of such matrices can create new opportunities for tissue engineering or provide a basis for optically stimulated drug depots. © 2019 WILEY-VCH Verlag GmbH & Co. KGaA, Weinheim

Ates, H. C. | Mohsenin, H. | Wenzel, C. | Glatz, R. T. | Wagner, H. J. | Bruch, R. | Hoefflin, N. | Spassov, S. | Streicher, L. | Lozano-Zahonero, S. | Flamm, B. | Trittler, R. | Hug, M. J. | Köhn, M. | Schmidt, J. | Schumann, S. | Urban, G. A. | Weber, Wilfried | Dincer, C.
DOI:
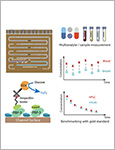
Personalized antibiotherapy ensures that the antibiotic concentration remains in the optimal therapeutic window to maximize efficacy, minimize side effects, and avoid the emergence of drug resistance due to insufficient dosing. However, such individualized schemes need frequent sampling to tailor the blood antibiotic concentrations. To optimally integrate therapeutic drug monitoring (TDM) into the clinical workflow, antibiotic levels can either be measured in blood using point-of-care testing (POCT), or can rely on noninvasive sampling. Here, a versatile biosensor with an antibody-free assay for on-site TDM is presented. The platform is evaluated with an animal study, where antibiotic concentrations are quantified in different matrices including whole blood, plasma, urine, saliva, and exhaled breath condensate (EBC). The clearance and the temporal evaluation of antibiotic levels in EBC and plasma are demonstrated. Influence of matrix effects on measured drug concentrations is determined by comparing the plasma levels with those in noninvasive samples. The system's potential for blood-based POCT is further illustrated by tracking ß‑lactam concentrations in untreated blood samples. Finally, multiplexing capabilities are explored successfully for multianalyte/sample analysis. By enabling a rapid, low-cost, sample-independent, and multiplexed on-site TDM, this system can shift the paradigm of “one‑size-fits-all” strategy. © 2021 The Authors. Advanced Materials published by Wiley-VCH GmbH
Mohsenin, Hasti | Wagner, Hanna J. | Rosenblatt, Marcus | Kemmer, Svenja | Dreppe, Friedel | Huesgen, Pitter | Timmer, Jens | Weber, Wilfried
DOI:
Synthetic biology applies concepts from electrical engineering and information processing to endow cells with computational functionality. Transferring the underlying molecular components into materials and wiring them according to topologies inspired by electronic circuit boards has yielded materials systems that perform selected computational operations. However, the limited functionality of available building blocks is restricting the implementation of advanced information-processing circuits into materials. Here, a set of protease-based biohybrid modules the bioactivity of which can either be induced or inhibited is engineered. Guided by a quantitative mathematical model and following a design-build-test-learn (DBTL) cycle, the modules are wired according to circuit topologies inspired by electronic signal decoders, a fundamental motif in information processing. A 2-input/4-output binary decoder for the detection of two small molecules in a material framework that can perform regulated outputs in form of distinct protease activities is designed. The here demonstrated smart material system is strongly modular and can be used for biomolecular information processing for example in advanced biosensing or drug delivery applications.

