Unsere Inspiration ist die Anpassungsfähigkeit von Organismen und den Materialien, aus denen sie aufgebaut sind, an wechselnde Umweltbedingungen. Pflanzen passen ihr Wachstum an die Lichtverhältnisse an, Bakterien entwickeln Resistenzen gegen Antibiotika oder Knochen werden durch Belastung stärker. Grundlage für diese Anpassungsfähigkeit ist eine faszinierende Signalverarbeitung der Organismen: Durch molekulare Sensoren werden Umweltbedingungen wahrgenommen, die Signale werden prozessiert und mit dem genetischen Programm des Organismus integriert, um am Ende eine passgenaue Reaktion auszulösen.
In unserer Forschung verwenden wir diese molekularen informationsverarbeitenden Mechanismen, um die Funktion und Eigenschaften von Zellen und Materialien gezielt zu steuern. Dies eröffnet neuartige Möglichkeiten in der grundladen- und anwendungsorientierten Forschung.
Mehr Informationen finden Sie auf unserer englischsprachigen Seite.

Mitarbeiter/innen
Publikationen
Mohsenin, Hasti | Pacheco, Jennifer | Kemmer, Svenja | Wagner, Hanna J. | Höfflin, Nico | Bergmann, Toquinha | Baumann, Tim | Jerez-Longres, Carolina | Ripp, Alexander | Jork, Nikolaus | Jessen, Henning J. | Fussenegger, Martin | Köhn, Maja | Timmer, Jens | Weber, Wilfried
DOI:
The site-specific and covalent conjugation of proteins on solid supports and in hydrogels is the basis for the synthesis of biohybrid materials offering broad applications. Current methods for conjugating proteins to desired targets are often challenging due to unspecific binding, unstable (noncovalent) coupling, or expensive and difficult-to-synthesize ligand molecules. Here, is presented PenTag, an approach for the bioorthogonal, highly specific, and covalent conjugation of a protein to its ligand for various applications in materials sciences. Penicillin-binding protein 3 (PBP3) is engineered and shows that this protein can be used for the stable and spontaneous conjugation of proteins to dyes, polymers, or solid supports. PenTag as a crosslinking tool is applied for synthesizing stimuli-responsive hydrogels or for the development of a biohybrid material system performing computational operations emulating a 4:2 encoder. Based on this broad applicability and the use of a small, cheap, and easy-to-functionalize ligand and a stable, soluble recombinant protein, is seen PenTag as a versatile approach toward biohybrid material synthesis.
Mohsenin, Hasti | Wagner, Hanna J. | Rosenblatt, Marcus | Kemmer, Svenja | Dreppe, Friedel | Huesgen, Pitter | Timmer, Jens | Weber, Wilfried
DOI:
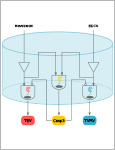
Synthetic biology applies concepts from electrical engineering and information processing to endow cells with computational functionality. Transferring the underlying molecular components into materials and wiring them according to topologies inspired by electronic circuit boards has yielded materials systems that perform selected computational operations. However, the limited functionality of available building blocks is restricting the implementation of advanced information-processing circuits into materials. Here, a set of protease-based biohybrid modules the bioactivity of which can either be induced or inhibited is engineered. Guided by a quantitative mathematical model and following a design-build-test-learn (DBTL) cycle, the modules are wired according to circuit topologies inspired by electronic signal decoders, a fundamental motif in information processing. A 2-input/4-output binary decoder for the detection of two small molecules in a material framework that can perform regulated outputs in form of distinct protease activities is designed. The here demonstrated smart material system is strongly modular and can be used for biomolecular information processing for example in advanced biosensing or drug delivery applications.
Armbruster, Anja | Mohamed, Asim M.E. | Phan, Hoang Trong | Weber, Wilfried
DOI:
Molecular optogenetics utilizes genetically encoded, light-responsive protein switches to control the function of molecular processes. Over the last two years, there have been notable advances in the development of novel optogenetic switches, their utilization in elucidating intricate signaling pathways, and their progress toward practical applications in biotechnological processes, material sciences, and therapeutic applications. In this review, we discuss these areas, offer insights into recent developments, and contemplate future directions.
Urban, Nadine | Hörner, Maximilian | Weber, Wilfried | Dincer, Can
DOI:
Circumventing the limitations of current bioassays, we introduce a light-controlled assay, OptoAssay, toward wash- and pump-free point-of-care diagnostics. Extending the capabilities of standard bioassays with light-dependent and reversible interaction of optogenetic switches, OptoAssays enable a bidirectional movement of assay components, only by changing the wavelength of light. Demonstrating exceptional versatility, the OptoAssay showcases its efficacy on various substrates, delivering a dynamic bioassay format. The applicability of the OptoAssay is successfully demonstrated by the calibration of a competitive model assay, resulting in a superior limit of detection of 8 pg ml−1, which is beyond those of conventional ELISA tests. In the future, combined with smartphones, OptoAssays could obviate the need for external flow control systems such as pumps or valves and signal readout devices, enabling on-site analysis in resource-limited settings.
Lohse, Stefan | Mink, Jan Niklas | Eckhart, Lea | Hans, Muriel Charlotte | Jusufi, Leuart | Zwick, Anabel | Mohr, Tobias | Bley, Isabelle Ariane | Khalmurzaev, Oybek | Matveev, Vsevolod Borisovich | Loertzer, Philine | Pryalukhin, Alexey | Hartmann, Arndt | Geppert, Carol-Immanuel | Loertzer, Hagen | Wunderlich, Heiko | Lehnhof, Hans-Peter | Naumann, Carsten Maik | Kalthoff, Holger | Junker, Kerstin
DOI:
PeCa is a rare entity with rising incidence rates due to increased infections with human papillomaviruses (HPV). The distinct subtypes of PeCa with an individual pathogenesis demand biomarkers for a precise patient risk assessment regarding disease progression and therapeutic susceptibility. We recently identified promising candidates associated with an HPV-instructed tumor microenvironment (TME) using HPV-positive PeCa cell lines and tissue microarrays (TMA). The capacity of HPV + p63 + PeCa cells to release neutrophil-attracting CXCL-8 provided a molecular link explaining the infiltration of CD15 + myeloid cells in PeCa specimens. The candidate biomarkers HPV, p63, CD15, DKK1, and CD147 linked a tumor-promoting TME with a higher TNM classification reflecting more aggressive and metastasizing cancers. Based on immune-reactive scores (IRS) from TMA staining for these biomarkers, we calculated correlations and conducted association analyses to assess the degree of relationship between all biomarkers. We then conducted Kaplan–Meier survival estimates and Cox regression analyses to delineate the impact on PeCa patient survival. There is a notable predictive potential regarding the survival of patients with biomarker profiles beyond the potency of the individual biomarker. From all candidate biomarkers and biomarker profiles, the combination of CD147 and infiltrating CD15 + cells linked to an active HPV-driven transformation displayed cancer-immune dynamics with dismal prognosis for patients. After deciphering relevant interdependencies, the HPV + CD147 + CD15 + status was the most potent profile predicting metastasis-free survival of PeCa patients. The results of this report underscore the need for analysis of the TME and the development of multi-parameter composite scores that reflect fundamental cancer-immune relationships to tailor therapeutic interventions based on actual cancer immune dynamics.
Valcenko, Alexander | Zwick, Anabel | Schneider, Lissy | Linxweiler, Maximilian | Lohse, Stefan
DOI:
Neutrophils play a crucial role in the tumor microenvironment (TME) of head and neck squamous cell carcinomas (HNSCC) and significantly influence treatment outcomes. Phenotypic and functional properties of neutrophils adapt to the TME with distinct subsets modulating disease progression and therapeutic interventions. Here, we evaluated phenotypic and functional differences of neutrophils derived from HNSCC patients and healthy donors. We observed significant phenotypic differences between neutrophils from healthy donors and HNSCC patient-derived neutrophils. Gender and tumor stage influenced neutrophil phenotypes and their ability to lyse tumor cells through antibody-dependent cell-mediated cytotoxicity (ADCC). Patients with advanced HNSCC and males may benefit less from neutrophil-centered immunotherapy. An engineered IgA2 antibody specific for the epidermal growth factor receptor (EGFR) demonstrated superior efficacy in activating neutrophils for ADCC compared to Panitumumab using healthy and patient-derived neutrophils, underscoring the potential of the IgA isotype as a therapeutic alternative. The distinct behavior and antibody-isotype dependent ADCC competence of CD177+/- neutrophils of healthy but not HNSCC donors warrants further exploration. Our study emphasizes the importance of personalized immunotherapy treatments that consider the characteristics of neutrophils, patient demographics, and the type of antibody to improve ADCC and ultimately enhance treatment outcomes for HNSCC.
Fischer, Alexandra A. M. | Schatz, Larissa | Baaske, Julia | Römer, Winfried | Weber, Wilfried | Thuenauer, Roland
DOI:
Each cell in a multicellular organism permanently adjusts the concentration of its cell
surface proteins. In particular, epithelial cells tightly control the number of carriers,
transporters and cell adhesion proteins at their plasma membrane. However, sensi-
tively measuring the cell surface concentration of a particular protein of interest in
live cells and in real time represents a considerable challenge. Here, we introduce a
novel approach based on split luciferases, which uses one luciferase fragment as a
tag on the protein of interest and the second fragment as a supplement to the extra-
cellular medium. Once the protein of interest arrives at the cell surface, the luciferase
fragments complement and generate luminescence. We compared the performance
of split Gaussia luciferase and split Nanoluciferase by using a system to synchronize
biosynthetic trafficking with conditional aggregation domains. The best results were
achieved with split Nanoluciferase, for which luminescence increased more than
6000-fold upon recombination. Furthermore, we showed that our approach can sep-
arately detect and quantify the arrival of membrane proteins at the apical and baso-
lateral plasma membrane in single polarized epithelial cells by detecting the
luminescence signals with a microscope, thus opening novel avenues for characteriz-
ing the variations in trafficking in individual epithelial cells.
Hörner, Maximilian | Becker, Jan | Bohnert, Rebecca | Banos, Miguel | Jerez-Longres, Carolina | Mühlhäuser, Vanessa | Härrer, Daniel | Wang Wong, Tin | Meier, Matthias | Weber, Wilfried
DOI:
Hydrogels with adjustable mechanical properties have been engineered as matrices for mammalian cells and allow the dynamic, mechano-responsive manipulation of cell fate and function. Recent research yields hydrogels, where biological photoreceptors translated optical signals into a reversible and adjustable change in hydrogel mechanics. While their initial application provides important insights into mechanobiology, broader implementation is limited by a small dynamic range of addressable stiffness. Herein, this limitation is overcome by developing a photoreceptor-based hydrogel with reversibly adjustable stiffness from ≈800 Pa to the sol state. The hydrogel is based on star-shaped polyethylene glycol, functionalized with the red/far-red light photoreceptor phytochrome B (PhyB), or phytochrome-interacting factor 6 (PIF6). Upon illumination with red light, PhyB heterodimerizes with PIF6, thus crosslinking the polymers and resulting in gelation. However, upon illumination with far-red light, the proteins dissociate and trigger a complete gel-to-sol transition. The hydrogel's light-responsive mechanical properties are comprehensively characterized and it is applied as a reversible extracellular matrix for the spatiotemporally controlled deposition of mammalian cells within a microfluidic chip. It is anticipated that this technology will open new avenues for the site- and time-specific positioning of cells and will contribute to overcome spatial restrictions.
Jerez-Logres, Carolina | Gómez-Matos, Marieta | Becker, Jan | Hörner, Maximilian | Wieland, Franz-Georg | Timmer, Jens | Weber, Wilfried
DOI:
Encapsulated cell-based therapies involve the use of genetically-modified cells embedded in a material in order to produce a therapeutic agent in a specific location in the patient's body. This approach has shown great potential in animal model systems for treating diseases such as type I diabetes or cancer, with selected approaches having been tested in clinical trials. Despite the promise shown by encapsulated cell therapy, though, there are safety concerns yet to be addressed, such as the escape of the engineered cells from the encapsulation material and the resulting production of therapeutic agents at uncontrolled sites in the body. For that reason, there is great interest in the implementation of safety switches that protect from those side effects. Here, we develop a material-genetic interface as safety switch for engineered mammalian cells embedded into hydrogels. Our switch allows the therapeutic cells to sense whether they are embedded in the hydrogel by means of a synthetic receptor and signaling cascade that link transgene expression to the presence of an intact embedding material. The system design is highly modular, allowing its flexible adaptation to other cell types and embedding materials. This autonomously acting switch constitutes an advantage over previously described safety switches, which rely on user-triggered signals to modulate activity or survival of the implanted cells. We envision that the concept developed here will advance the safety of cell therapies and facilitate their translation to clinical evaluation.
Molinari, Pamela E. | Krapp, Adriana R. | Weiner, Andrea | Beyer, Hannes M. | Kondadi, Arun Kumar | Blomeier, Tim | López, Melina | Bustos-Sanmamed, Pilar | Tevere, Evelyn | Weber, Wilfried | Reichert, Andreas S. | Calcaterra, Nora B. | Beller, Mathias | Carrillo, Nestor | Zurbriggen, Matias D.
DOI:
NADP(H) is a central metabolic hub providing reducing equivalents to multiple biosynthetic, regulatory and antioxidative pathways in all living organisms. While biosensors are available to determine NADP+ or NADPH levels in vivo, no probe exists to estimate the NADP(H) redox status, a determinant of the cell energy availability. We describe herein the design and characterization of a genetically-encoded ratiometric biosensor, termed NERNST, able to interact with NADP(H) and estimate ENADP(H). NERNST consists of a redox-sensitive green fluorescent protein (roGFP2) fused to an NADPH-thioredoxin reductase C module which selectively monitors NADP(H) redox states via oxido-reduction of the roGFP2 moiety. NERNST is functional in bacterial, plant and animal cells, and organelles such as chloroplasts and mitochondria. Using NERNST, we monitor NADP(H) dynamics during bacterial growth, environmental stresses in plants, metabolic challenges to mammalian cells, and wounding in zebrafish. NERNST estimates the NADP(H) redox poise in living organisms, with various potential applications in biochemical, biotechnological and biomedical research.

