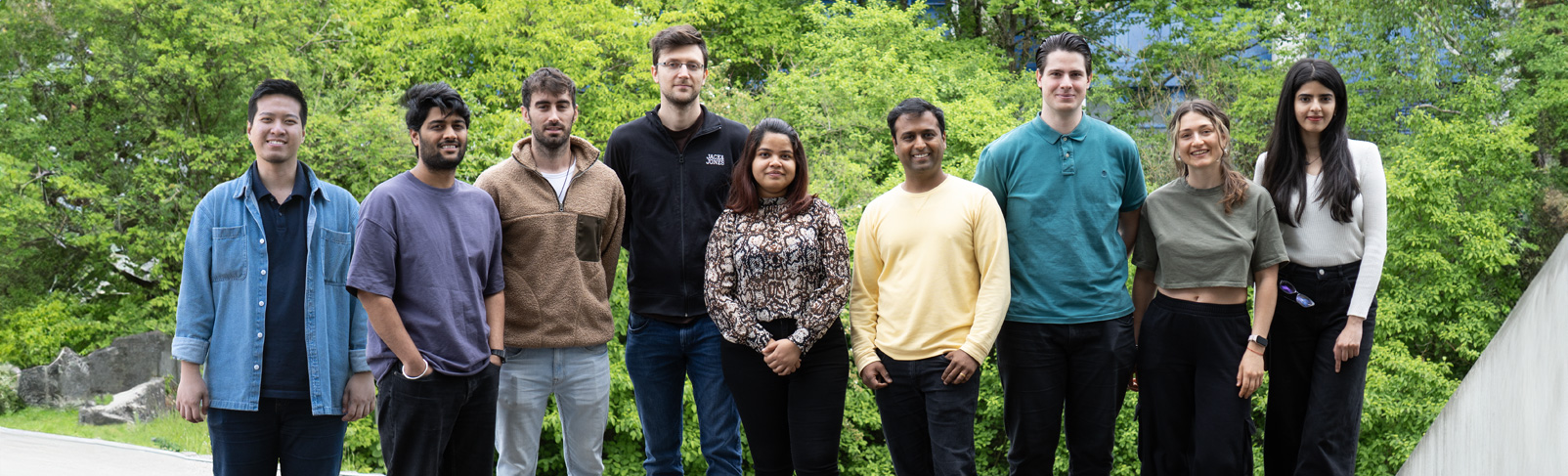
Die Forschungsgruppe Bioprogrammierbare Materialien erforscht ein junges multidisziplinäres Feld, das synthetische Biologie und Biomaterialien kombiniert. Im Mittelpunkt steht die Entwicklung von Materialien mit genetisch programmierten Funktionalitäten, die für Biosensorik geeignet sowie in der Lage sind, auf Stimuli hin Medikamente langfristig freizusetzen und das Zellverhalten zu beeinflussen. Mit Werkzeugen aus der synthetischen Biologie werden Proteine und Mikroben so programmiert, dass sie intelligente und nützliche Funktionen erfüllen. Diese technisierten biologischen Gebilde werden dann in entsprechend entwickelte polymere Matrizen eingearbeitet. Das Ergebnis sind Kompositmaterialien mit sehr vielseitigen Funktionalitäten, einer großen Bandbreite an Steuerbarkeit und in situ-Kontrollmöglichkeiten.
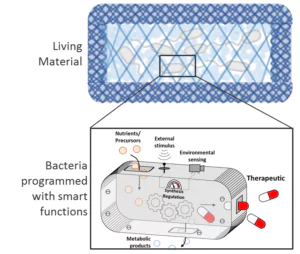
Bakterien werden mit intelligenten Funktionen versehen, beispielsweise für Anwendungen in den Bereichen Biosensorik und Wirkstofftransport

Mitarbeiter/innen





Forschung
In der Gruppe Bioprogrammierbare Materialien kombinieren wir genetisch programmierte lebende Organismen wie Bakterien mit polymeren Matrices wie Hydrogelen, um intelligente Verbundwerkstoffe, so genannte Engineered Living Materials (ELMs), herzustellen. Durch innovative Ansätze bei den lebenden wie bei den nicht-lebenden Materialien sind unsere ELMs für ganze eine Reihe von biomedizinischen Anwendungen wie Biosensorik und Medikamentenabgabe geeignet:
Stimuli-responsive Drug-Sekretion in Bakterien
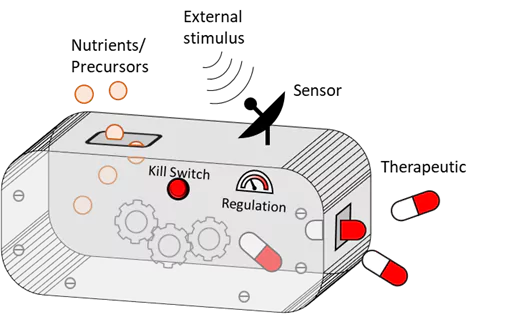
Bakterien sind ein wesentlicher Bestandteil des Mikrobioms des Körpers, wobei mehrere native und probiotische Arten dem Menschen einen natürlichen Gesundheitsschutz verleihen. Bakterien werden auch in der pharmazeutischen Industrie in großem Umfang als Biofabriken zur Herstellung einer Vielzahl von Arzneimitteln eingesetzt. Unsere Forschung vereint diese beiden Merkmale: Wir entwickeln medizinisch relevante Bakterien wie E. coli, Lactobacilli und Corynebacterium, um therapeutische Moleküle direkt im Körper dort zu produzieren und abzusondern, wo sie benötigt werden. Da die Bakterien auf natürliche Weise im Körper gedeihen, kann eine langfristige Freisetzung des Arzneimittels aufrechterhalten werden. Wir designen die Bakterien so, dass sie antimikrobielle, entzündungshemmende und regenerative Medikamente in Form von Peptiden, Proteinen und enzymatisch synthetisierten Biomolekülen zur Behandlung chronischer Krankheiten produzieren und freisetzen.
Patientenprofile und Krankheitsverläufe können sehr unterschiedlich sein. Daher sollte die Aktivität dieser therapeutischen Bakterien so personalisiert und angepasst werden können, dass eine größtmögliche Wirksamkeit sichergestellt ist. Dazu entwickeln wir genetische Module, die die Wirkstofffreisetzung über externe Reize„anschalten“. Stimuli wie Licht, Wärme und kleine Moleküle ermöglichen die Fernsteuerung der Bakterien, und Stimuli wie Biomarker reagieren auf das Fortschreiten der Krankheit und können die Aktivität der Bakterien mittels Autoregulation verändern. Stimuli-responsive genetische Module werden ebenfalls als zusätzliche Ebene der Biosicherheit entwickelt. Damit stellen wir sicher, dass Bakterien nicht außerhalb ihrer vorgesehenen Implantatumgebung überleben. Während viele der von uns verwendeten genetischen Module ursprünglich für E. coli entwickelt wurden, statten wir nun auch probiotische Bakterien wie Laktobazillen und Corynebacterium mit ähnlichen Funktionen aus – eine Herausforderung, die durch die Erweiterung des Spektrums an ELM-fähigen Organismen und Funktionen erhebliche Vorteile verspricht.
Neuste Veröffentlichungen:
- Dhakane, P.; Tadimarri, V. S.; Sankaran, S. Light-Regulated pro-Angiogenic Engineered Living Materials. bioRxiv – https://doi.org/10.1101/2022.10.28.514190.
- Dey, S.; Asensio, M. B.; Kuttae, S. B.; Sankaran, S. Novel Genetic Modules Encoding High-Level Antibiotic-Free Protein Expression in Probiotic Lactobacilli. bioRxiv – https://doi.org/10.1101/2022.08.04.502766.
Hydrogele für die bakterielle Einkapselung und Werkzeuge zum Verständnis des bakteriellen Verhaltens in Gefangenschaft
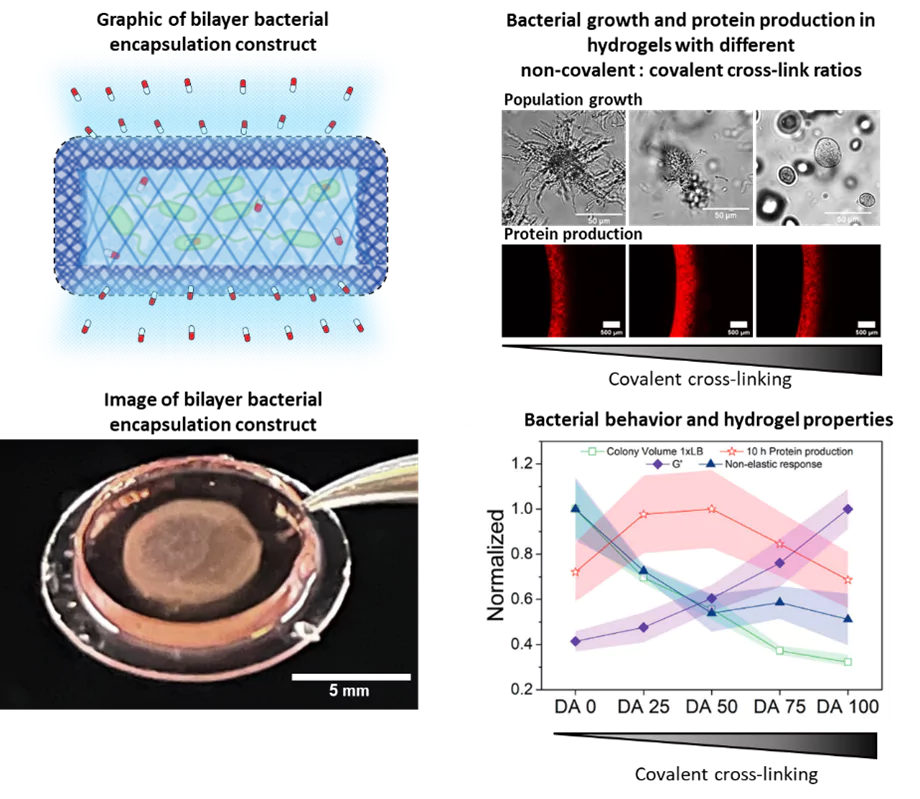
Für die therapeutischen Bakterien zur Behandlung einer Krankheit ist es notwendig, dass sie sich an der Krankheitsstelle ansiedeln und eine ausreichend große Population aufbauen, um wirksame Medikamentendosen bereitzustellen. In Zusammenarbeit mit der Forschungsgruppe Dynamische Biomaterialien entwickeln wir hydrogelbasierte Matrizen, die geeignete Umgebungen für das Wachstum und die Funktion der Bakterien bieten. Diese Hydrogele sind so konzipiert, dass sie die Größe der enthaltenen Bakterienpopulation mechanisch kontrollieren und gleichzeitig deren Funktionalität maximieren. Das Hydrogelnetzwerk ermöglicht die Diffusion von Nährstoffen, Metaboliten und Medikamenten in und aus der Matrix, während es das Entweichen der Bakterien verhindert und Schutz vor Immunzellen bietet. Die Materialkomponente bietet daher eine zusätzliche Ebene der Biosicherheit für den Einsatz von genetisch veränderten Bakterien im Körper.
Im Rahmen dieser Forschung haben wir festgestellt, dass bestimmte mechanische Eigenschaften der Hydrogele das Wachstum und den Stoffwechsel der enthaltenen Bakterien beeinflussen, ähnlich wie es bei natürlichen Biofilmen der Fall ist. Dies wirkt sich wiederum auf die Leistungsfähigkeit der Bakterien in Bezug auf die Reaktion auf Stimuli und die Produktion von Medikamenten aus. Daher untersuchen wir das Verhalten der Bakterien in den eingeschränkten Bedingungen der Hydrogele, deren mechanische Eigenschaften wir steuern können. Dabei erkennen wir das Zusammenspiel zwischen dem kollektiven Verhalten der Bakterien und den sie umgebenden mechanischen Kräften. Neben der Optimierung der Leistungsfähigkeit der von uns entwickelten lebenden Materialien dient dies auch als künstliche Plattform für Biofilme, um grundlegende Erkenntnisse über das mögliche Verhalten von Bakterien in verschiedenen Bereichen natürlicher Biofilme zu gewinnen. Zur Bestimmung des Wachstums und des Stoffwechselverhaltens der Bakterien im Laufe der Zeit nutzen wir Mikroskopie sowie biochemische und genetische Tests.
Neuste Veröffentlichungen:
- Bhusari, S.; Sankaran, S.; del Campo, A. Regulating Bacterial Behavior within Hydrogels of Tunable Viscoelasticity. Advanced Science 2022, 9 (17), 2106026. https://doi.org/10.1002/advs.202106026.
- Bhusari, S.; Kim, J.; Polizzi, K.; Sankaran, S.; Campo, A. del. Encapsulation of Bacteria in Bilayer Pluronic Thin Film Hydrogels: A Safe Format for Engineered Living Materials. bioRxiv – https://doi.org/10.1101/2022.09.29.510162.
Kooperationen:
Diese Forschungsanstrengungen sind auch Teil mehrerer Kooperationen innerhalb zweier Konsortien – (i) Leibniz Wissenschaftscampus – Living Therapeutic Materials (LSC LifeMat) und (ii) Sonderforschungsbereich on Physikalische Modellierung von Nichtgleichgewichtsprozessen in biologischen Systemen (SFB 1027).
Finanzierung:


Publikationen
Tadimarri, Varun S. | Blanch-Asensio, Marc | Deshpande, Ketaki | Baumann, Jonas | Baumann, Carole | Müller, Rolf | Trujillo, Sara | Sankaran, Shrikrishnan
DOI:
Engineered living materials (ELMs) made of bacteria in hydrogels have shown considerable promise for therapeutic applications through controlled and sustained release of complex biopharmaceuticals at low costs and with reduced wastage. While most therapeutic ELMs use E. coli due to its large genetic toolbox, most live biotherapeutic bacteria in development are lactic acid bacteria due to native health benefits they offer. Among these, lactobacilli form the largest family of probiotics with therapeutic potential in almost all sites of the body with a microbiome. A major factor limiting the use of lactobacilli in ELMs is their limited genetic toolbox. This study expands on recent work to expand the genetic programmability of probiotic Lactiplantibacillus plantarum WCFS1 for protein secretion and encapsulate it in a simple, cost-effective, and biocompatible core–shell alginate bead to develop an ELM. The controlled release of recombinant proteins is demonstrated, even up to 14 days from this ELM, thereby terming it PEARL – Protein Eluting Alginate with Recombinant Lactobacilli. Notably, lactobacillus encapsulation offered benefits like bacterial containment, protein release profile stabilization, and metabolite-induced cytotoxicity prevention. These findings demonstrate the mutual benefits of combining recombinant lactobacilli with alginate for the controlled and sustained release of proteins.
Dupont, L. | Tadimarri, Varun S. | Buret, R. | Sankaran, Shrikrishnan | Picton, L. | Jonas, A.M. | Glinel, K.
DOI:
Engineered living materials (ELMs) rely on the ability to control cell behavior in material systems. ELMs containing bacteria secreting beneficial molecules are being developed for therapeutic purposes. Using commensal strains embedded in physically cross-linked agarose hydrogels, we systematically investigate how gel rigidity and initial bacterial density affect the morphology of bacterial colonies and their secretory function. Although often considered independently, these parameters jointly define the microscale environment experienced by embedded cells, influencing nutrient access, mechanical interactions, and potential cell-to-cell communication. We show that matrix rigidity effectively tunes aggregate morphology, modulating their shape and compactness, without compromising bacterial growth or secretion. In parallel, initial bacterial density determines the biomass accumulation dynamics and spatial distribution of aggregates, which in turn influence the onset and temporal profile of secretory activity, without altering its final magnitude. This decoupling between structural organization and secretory output opens new possibilities for engineering ELMs with tailored architectures and prolonged secretory and release activity.
Desai, Krupansh K. | Sankaran, Shrikrishnan | Del Campo, Aránzazu | Trujillo, Sara
DOI:
Engineered living materials (ELMs), which usually comprise bacteria, fungi, or animal cells entrapped in polymeric matrices, offer limitless possibilities in fields like drug delivery or biosensing. Determining the conditions that sustain ELM performance while ensuring compatibility with ELM hosts is essential before testing them in vivo. This is critical to reduce animal experimentation and can be achieved through in vitro investigations. Currently, there are no standards that ensure ELM compatibility with host tissues. Towards this goal, we designed a 96-well plate-based screening method to streamline ELM growth across culture conditions and determine their compatibility potential in vitro. We showed proliferation of three bacterial species encapsulated in hydrogels over time and screened six different cell culture media. We fabricated ELMs in bilayer and monolayer formats and tracked bacterial leakage as a measure of ELM biocontainment. After screening, an appropriate medium was selected that sustained growth of an ELM, and it was used to study cytocompatibility in vitro. ELM cytotoxicity on murine fibroblasts and human monocytes was studied by adding ELM supernatants and measuring cell membrane integrity and live/dead staining, respectively, proving ELM cytocompatibility. Our work illustrates a simple setup to streamline the screening of compatible environmental conditions of ELMs with the host.
Khazem, Ali | Schmachtenberg, Rosanne | Weiand, Anke | Sankaran, Shrikrishnan | Weber, Wilfried
DOI:
Living therapeutic and diagnostic materials based on engineered microorganisms are emerging as a novel approach with the perspective of providing patient-tailored, sustainable, and cost-effective healthcare solutions. In this review, we focus on recent advances in using genetically or chemically engineered microorganisms as living diagnostics, therapeutics, and as a means of prevention for various diseases. We also highlight the applications of living therapeutics for acute and chronic diseases, and the role of micro/macro-encapsulation of the engineered microorganisms. We further showcase the current success of engineered living therapeutics in clinical trials and discuss challenges and future trends in the field.
Tadimarri, Varun Sai | Tyagi, Tanya Amit | Duong, Cao Nguyen | Rasheed, Sari | Müller, Rolf | Sankaran, Shrikrishnan
DOI:
Encapsulation of microbes in natural or synthetic matrices is a key aspect of engineered living materials, although the influence of such confinement on microbial behavior is poorly understood. A few recent studies have shown that the spatial confinement and mechanical properties of the encapsulating material significantly influence microbial behavior, including growth, metabolism, and gene expression. However, comparative studies within different bacterial species under identical confinement conditions are limited. In this study, Gram-negative Escherichia coli Nissle 1917 and Gram-positive Lactiplantibacillus plantarum WCFS1 were encapsulated in hydrogel matrices, and their growth, metabolic activity, and recombinant gene expression were examined under varying degrees of hydrogel stiffness, achieved by adjusting the polymer concentration and chemical cross-linking. Both bacteria grow from single cells into confined colonies, but more interestingly, in E. coli gels, mechanical properties influenced colony growth, size, and morphology, whereas this did not occur in L. plantarum gels. However, with both bacteria, increased matrix stiffness led to higher levels of recombinant protein production within the colonies. By measuring metabolic heat from the bacterial gels using the isothermal microcalorimetry technique, it was inferred that E. coli adapts to the mechanical restrictions through multiple metabolic transitions and is significantly affected by the different hydrogel properties. Contrastingly, both of these aspects were not observed with L. plantarum. These results revealed that despite both bacteria being gut-adapted probiotics with similar geometries, mechanical confinement affects them considerably differently. The weaker influence of matrix stiffness on L. plantarum is attributed to its slower growth and thicker cell wall, possibly enabling the generation of higher turgor pressures to overcome restrictive forces under confinement. By providing fundamental insights into the interplay between mechanical forces and bacterial physiology, this work advances our understanding of how matrix properties shape bacterial behavior. The implications of these findings will aid the design of engineered living materials for therapeutic applications.
Deshpande, Ketaki | Tadimarri, Varun S. | Ramirez-Rangel, Juliette | Sankaran, Shrikrishnan | Trujillo, Sara
DOI:
Living therapeutics are attractive candidates to tackle the limitations of classically delivered therapeutic peptides, which are often poorly stable and require cost-intensive modifications. Their functional assessment is limited to animal experiments, which increase the complexity to evaluate the dynamic nature of these systems. Therefore, we developed an in vitro model of endotoxemia using macrophages to assess early-stage anti-inflammatory Living therapeutics. We refined the model based on three anti-inflammatory peptides (KCF-18, I6P7, and α-MSH) and identified suitable therapeutic concentrations and treatment durations. We applied the model to Lactiplantibacillus plantarum TF103, a probiotic engineered to secrete these peptides. The model revealed that Living therapeutics enhanced the effects of the peptides, requiring lower amounts of anti-inflammatory effects. This points to potential synergistic effects between peptides and bacteria. The model presented here allows the investigation of dynamic regimes, which could be useful in the development of complex systems such as the ones encountered in Living therapeutics.
Blanch-Asensio, Marc | Tadimarri, Varun S. | Martinez, Roberto | Singh Dahiya, Gurvinder | Duong, Cao Nguyen | Lale, Rahmi | Sankaran, Shrikrishnan
DOI:
Living microbial therapeutics promise precise, programmable interventions at disease sites, yet most demonstrations of on demand drug release still rely on Escherichia coli, whose rich genetic toolkit is unmatched among probiotics. In particular, genetic parts to regulate in situ protein production are severely lacking in non-model probiotic bacteria like lactobacilli. Here, we equip the probiotic Lactiplantibacillus plantarum with high-performance genetic switches and show how material encapsulation can further enhance their behavior. By integrating cumate or vanillate-responsive operators and repressors with the strongest constitutive promoter in L. plantarum (Ptec), we generated two switches that support micromolar range induction. In rapidly growing culture conditions, acidification-associated leakiness of the switch was observed, which could compromise their applicability for precise on-demand delivery of drugs. Furthermore, such leakiness also limits the duration for which these engineered probiotics can be reliably used. By restricting growth through mild temperature or nutrient limitation, acidification and leakiness were suppressed. Strikingly, immobilizing the engineered cells in core-shell alginate beads (Protein Eluting Alginate with Recombinant Lactobacilli, PEARLs) almost eliminated leakiness, enabling day-scale, reversible control of intracellular reporters and secreted enzymes. This leakiness suppression persisted when two strains carrying orthogonal switches were co-encapsulated and even after miniaturization to submillimeter beads. These results expand the genetic toolbox of probiotic L. plantarum, demonstrate the synergy between genetic circuit design and material encapsulation, and advance lactobacilli toward stimuli-responsive therapeutic platforms.
Bhusari, Shardul | Hoffmann, Maxi | Herbeck-Engel, Petra | Sankaran, Shrikrishnan | Wilhelm, Manfred | del Campo, Aránzazu
DOI:
Pluronic (Plu) hydrogels mixed with variable fractions of Pluronic diacrylate (PluDA) have become popular matrices to encapsulate bacteria and control their growth in engineered living materials. Here we study the rheological response of 30 wt.% Plu/PluDA hydrogels with PluDA fraction between 0 and 1. We quantify the range of viscoelastic properties that can be covered in this system by varying in the PluDA fraction. We present stress relaxation and creep-recovery experiments and describe the variation of the critical yield strain/stress, relaxation and recovery parameters of Plu/PluDA hydrogels as function of the covalent crosslinking degree using the Burgers and Weilbull models. The analyzed hydrogels present two stress relaxations with different timescales which can be tuned with the covalent crosslinking degree. We expect this study to help users of Plu/PluDA hydrogels to estimate the mechanical properties of their systems, and to correlate them with the behaviour of bacteria in future Plu/PluDA devices of similar composition.
Blanch-Asensio, Marc | Tadimarri, Varun S. | Wilk, Alina | Sankaran, Shrikrishnan
DOI:
Background: The Lactobacillaceae family comprises many species of great importance for the food and healthcare industries, with numerous strains identified as beneficial for humans and used as probiotics. Hence, there is a growing interest in engineering these probiotic bacteria as live biotherapeutics for animals and humans. However, the genetic parts needed to regulate gene expression in these bacteria remain limited compared to model bacteria like E. coli or B. subtilis. To address this deficit, in this study, we selected and tested several bacteriophage-derived genetic parts with the potential to regulate transcription in lactobacilli.
Results: We screened genetic parts from 6 different lactobacilli-infecting phages and identified one promoter/repressor system with unprecedented functionality in Lactiplantibacillus plantarum WCFS1. The phage-derived promoter was found to achieve expression levels nearly 9-fold higher than the previously reported strongest promoter in this strain and the repressor was able to almost completely repress this expression by reducing it nearly 500-fold.
Conclusions: The new parts and insights gained from their engineering will enhance the genetic programmability of lactobacilli for healthcare and industrial applications.
Blanch Asensio, Marc | Dey, Sourik | Tadimarri, Varun S. | Sankaran, Shrikrishnan
DOI:
Lactobacilli are ubiquitous in nature and symbiotically provide health benefits for countless organisms including humans, animals and plants. They are vital for the fermented food industry and are being extensively explored for healthcare applications. For all these reasons, there is considerable interest in enhancing and controlling their capabilities through the engineering of genetic modules and circuits. One of the most robust and reliable microbial chassis for these synthetic biology applications is the widely used Lactiplantibacillus plantarum species. However, the genetic toolkit needed to advance its applicability remains poorly equipped. This mini-review highlights the genetic parts that have been discovered to achieve food-grade recombinant protein production and speculates on lessons learned from these studies for L. plantarum engineering. Furthermore, strategies to identify, create and optimize genetic parts for real-time regulation of gene expression and enhancement of biosafety are also suggested.




