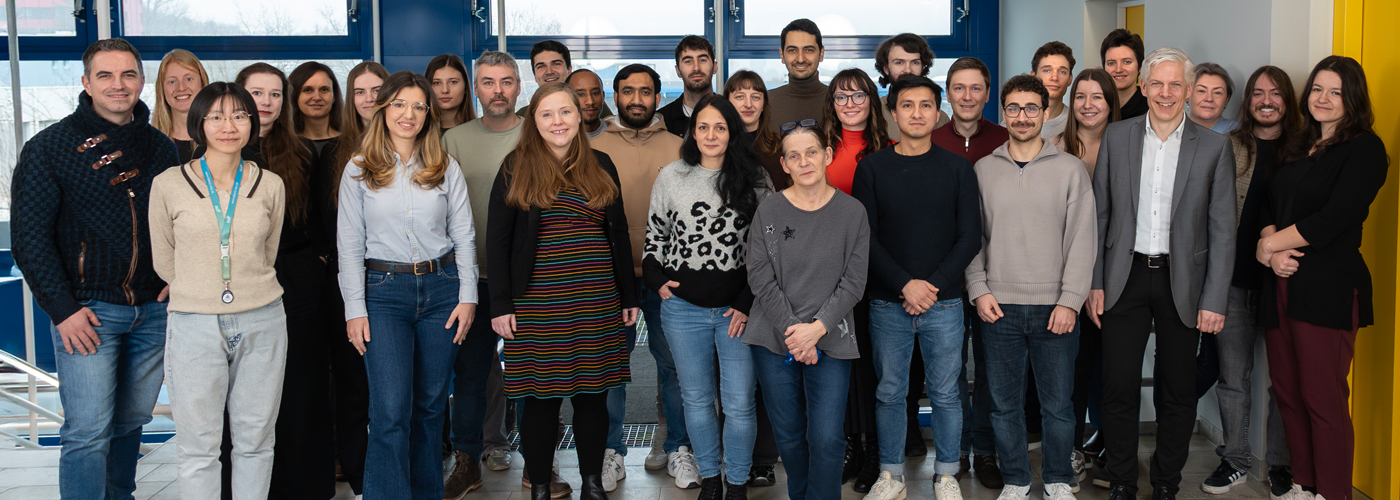
We engineer cells and materials that communicate and process information through synthetic biology
Our inspiration is the ability of organisms and the materials they are made of to adapt to dynamic environmental conditions. Plants adapt growth to light conditions; bacteria develop resistance against antibiotics or bones get stronger when exercised. The basis for this ability to adapt is a fascinating information processing machinery of the organisms: Environmental conditions are captured by molecular sensors, then the signals are processed and integrated with genetic programs to finally yield a targeted response.
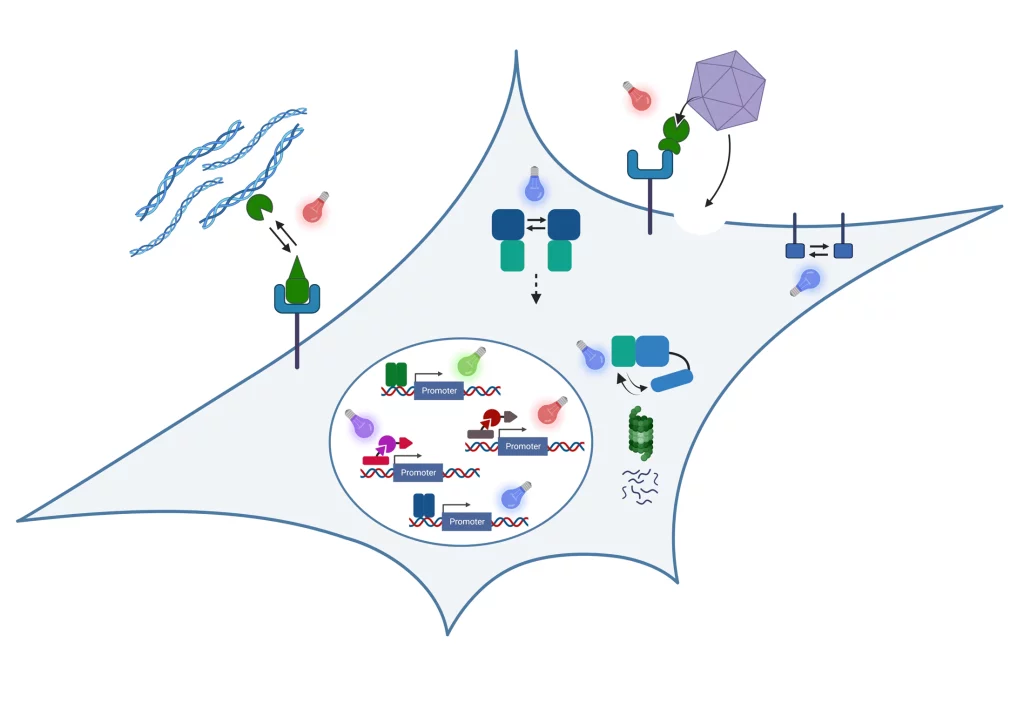
In our research, we engineer nature’s molecular sensing, processing, and actuation machinery in order to precisely control the function and properties of cells and materials. We apply these newly developed technologies in different fields of fundamental and applied research.

Team Members











Research
Stimulus-responsive and Information-processing (living) Materials

We develop and apply stimulus-responsive and information-processing biohybrid polymer materials. To this aim, we functionally couple synthetic biological molecular sensors and switches to polymer materials. By wiring these switches according to topologies inspired by electronic circuits, we engineer materials that perform fundamental computational operations. Examples of our work include:
- We engineered a hydrogel based on a bacteria-derived photoreceptor which allows the light-responsive, fully reversibly tuning of its mechanical properties. We applied this hydrogel as extracellular matrix to analyze the impact of dynamic mechanical environments on transcriptome-wide responses in mesenchymal stem cells or on the migration of T-lymphocytes.
See Hörner et al. Advanced Materials 2019 - We integrated synthetic biological switches with polymer materials into a circuit inspired by an electronic counter. The resulting material system was able to count the number of input light pulses and to release different output as a function of the number of light pulses detected. We applied this system to sequentially release different biocatalysts to drive a two-step biochemical reaction.
See Beyer et al., Advanced Materials 2018 - We developed PenTag, a protein tag for the spontaneous, covalent coupling of proteins to ampicillin-functionalized molecules such as dyes, polymers, or solid supports. Based on this strategy, we engineered and assembled material modules to function as encoder for processing different combinations of biochemical input stimuli.
See Mohsenin et al., Advanced Functional Materials 2024 - By engineering modular protease-based switches that can either be activated or repressed, we develop information-processing biohybrid circuits that process binary biomolecular information according to a circuit inspired by electronic decoders. Such circuits can be applied to process and interpret biochemical sensor information for advanced diagnostic applications.
See Mohsenin et al., Advanced Materials 2024
Molecular optogenetics to control cell fate and function
We develop and apply molecular optogenetic tools to control cell fate and function with unprecedented spatial and temporal precision in a dose-dependent and highly specific manner. To this aim, we engineer plant- and bacteria-derived photoreceptors and functionally couple them to proteins involved in cell signaling and gene expression. Examples of our work include:
- Light-inducible formation of liquid or gel-like transcription factor condensates in mammalian cells and mice. We demonstrate that liquid “transcription factor droplets” show a several-fold higher activity in inducing transgene expression compared to native transcription factors. Further, gel-like transcription factor condensates were shown to correlate with decreased transcriptional activation thus providing a materials-based layer of controlling gene expression.
See Schneider et al., Science Advances 2021 and Fischer et al., Small 2024 - Light-guided adeno-associated viral (AAV) vectors. We engineered a light-responsive tropism into AAVs which allows us to selectively transfer genetic information into single cells or to transduce different cells within one culture with different transgenes.
See Hörner et al., Science Advances 2021
Our group is running www.optobase.org, the most comprehensive database on molecular optogenetics. Have a look and discover the amazing opportunities in controlling biology with light!
Biosensors
We integrate natural and engineered molecular sensors for drugs, metabolites or nucleic acids into suitable readout formats for the fast and sensitive quantification of such substances. Together with collaboration partners, we develop biosensor systems for different application fields:
Open Positions
We are always excited to meet curious and creative scientists passionate about synthetic biology, optogenetics, and engineered living materials. If you would like to shape the future of biobased and living materials with us, we warmly welcome your spontaneous application for a PhD thesis or Postdoc position!
Projects and Partners
We perform collaborative research in materials-oriented synthetic biology within interdisciplinary research consortia
STEADY
Within the ERC Advanced Grant STEADY, we develop concepts for dynamically controlling the properties of engineered living materials by advanced synthetic genetic circuits.
LoopOfFun
We coordinate the European Innovation Council (EIC)-funded consortium LoopOfFun in which we aim at developing a platform for the rapid development of industry-scale, one-step, simple casting-based manufacturing processes for fungal mycelia composites. We jointly work towards this goal with our consortium partners:
- Prof. Roman Jerala, National Institute of Chemistry, Ljubljana, Slovenia
- Dr. Achim Weber, Fraunhofer IGB, Stuttgart, Germany
- Prof. Arnold Driessen, University of Groningen, The Netherlands
- Carlotta Borgato and Jan Boelen, Atelier LUMA, Arles, France
DELIVER
In the project DELIVER funded by the Carl-Zeiss-Foundation, we collaborate towards the data-driven engineering of sustainable living materials. We combine synthetic biology with materials sciences and data-driven approaches to design bio-based composite materials with custom-tailored structural properties for construction applications. Within deliver, we collaborate with the following partners:
- Prof. Thomas Speck, University of Freiburg, Germany
- Dr. Clemens Kreutz, University Hospital Freiburg, Germany
BILLARD
We coordinate the BILLARD project funded by the Federal Ministry of Education and Research (BMBF) within the funding line “Biologization of Technology”, we collaborate with PD Dr. Felicitas Bucher from the Clinic of Ophtamology at the University Hospital Freiburg on the development of novel intraocular drug delivery devices.
CIBSS – Centre for Integrative Biological Signalling Studies
We are member of the Cluster of Excellence CIBSS in which we perform research on novel optogenetic technologies to control signaling reactions in mammalian cells. We mainly collaborate with Prof. Dr. Jens Timmer on the model-based design of synthetic biological switches and networks and with Prof. Dr. Wolfgang Schamel on controlling immunological processes such as T cell activation via optogenetics.
Publications
Heilmann, Heiko | Busch, Lukas | Buchmann, Celine | Mohamed, Islam | Theiß, Adrian | Junkger, Sabryna | Lohse, Stefan | Bufe, Bernd
DOI:
Formyl peptide receptors (FPRs) are pattern recognition receptors well-known for bacterial pathogen sensing. We here identified activator and inhibitor motifs for FPRs that are present on surface proteins of various viral pathogens. Peptides containing these motifs interact with all FPR family members and modulate various important immune functions in innate immune cells. Viral breakdown products comprising these motifs were found in patients with COVID-19. In the spike protein, many activators are found in highly mutagenic regions, whereas the inhibitor motif is located in a conserved domain that also exists in further unrelated viruses. The physiochemical properties of FPR1 activators correlate with the occurrence of protein aggregation hotspots. Such hotspots are present on various surface proteins of unrelated viruses that can also activate FPRs. This points toward a general contribution of FPRs in modulating antiviral immune responses during many distinct viral infections.
Leydet, Létitia | Couillaud, Julie | Amouric, Agnès | Courvoisier-Dezord, Elise | Avesque, Carole | Giardina, Thierry | Attolini, Mireille | Rousselot-Pailley, Pierre | Duquesne, Katia | Rosso, Marie-Noelle | Iacazio, Gilles
DOI:
Long-lasting polypore fungi are significant producers of terpene cyclases of high interest for medicinal or biotechnological applications. Following the 1000 Fungal Genomes initiative launched by the Joint Genome Institute, the genome of Cubamyces (C.) menziesii and identified 18 genes encoding sesquiterpene cyclases (STCs) is explored. In a search for robust catalysts suitable for practical applications, the 18 codon-optimized open reading frames are cloned and overproduced the C. menziesii STCs in Escherichia coli. In ten cases, the catalytically active enzyme is purified and tested with three chemically synthesized linear diphosphates: geranyl diphosphate, farnesyl diphosphate (FDP), and geranylgeranyl diphosphate. Only FDP proved to be a substrate for these 10 enzymes. The product specificity of all these enzymes is determined by (GC-MS) gas chromatography mass spectrometry and (NMR) nuclear magnetic resonance analysis. Among the 10 enzymes, four produced a predominant compound, four yielded two main compounds, and the remaining two acted as a multiproduct catalysts. This work sheds light on the potential sesquiterpenes involved in the chemical ecology of the polypore C. menziesii and provides evidence for the potential of Polyporales fungi in the identification of new sesquiterpene cyclase activities.
Greco, Adriana | Masselli, Claudia | Orlu, Mine | Weber, Wilfried
DOI:
Elastocaloric technology is a new way to heat and cool spaces by using stretchy metals, called shape-memory alloys, instead of harmful refrigerant gases. When these metals are squeezed or stretched, they heat up; and when they relax, they cool down. This process is called the elastocaloric effect and it is more energy efficient than traditional cooling systems, making it a cleaner, greener alternative. Elastocaloric systems could cool homes, schools, and workplaces, and they could refrigerate food and medicine in areas with limited electricity. Researchers are also testing this technology for cooling and heating of electric vehicles, where it could help conserve battery life, and for heating buildings in colder climates. Despite its promise, elastocaloric technology faces challenges, such as improving the durability of materials and making the shape-memory alloys more affordable. With continued research, this technology could someday help to reduce greenhouse gas emissions, lower energy costs, and bring life-saving cooling to more people all over the world.
Armbruster, Anja | Hörner, maximilian | Weber, Wilfried
DOI:
Methods for the precise temporal control of cell surface receptor activation are indispensable for the investigation of signaling processes in mammalian cells. Optogenetics enables such precise control, but its application in primary cells is limited by the imperative for genetic manipulation of target cells. We here describe a method that overcomes this obstacle and enables the precise activation of the T cell receptor in nongenetically engineered human T cells by light. Our optogenetic receptor activation system OptoREACT employs a TCR-specific scFv fused to PIF6 that interacts with tetramerized PhyB in a light-dependent manner and thereby clusters and activates the T cell receptor in response to red light. OptoREACT not only omits genetic manipulation of the target cell but, because of its modular nature, is likely applicable to a broad range of oligomerization-activated cell surface receptors.
Schmachtenberg, Rosanne | Weber, Wilfried
DOI:
The construction and assembly of information-processing biomaterials are limited by the need for laborious assembly of various circuits. A new framework to assemble protein-based elements encoding complex Boolean operations enables user-defined release of biomolecules from these materials.
Fischer, Alexandra A. M. | Kramer, Markus M. | Banos, Miguel | Grimm, Merlin M. | Fliegauf, Manfred | Grimbacher, Bodo | Radziwill,Gerald | Rahmann, Sven | Weber, Wilfried
DOI:
Molecular optogenetics allows the control of molecular signaling pathways in response to light. This enables the analysis of the kinetics of signal activation and propagation in a spatially and temporally resolved manner. A key strategy for such control is the light-inducible clustering of signaling molecules, which leads to their activation and subsequent downstream signaling. In this work, an optogenetic approach is developed for inducing graded clustering of different proteins that are fused to eGFP, a widely used protein tag. To this aim, an eGFP-specific nanobody is fused to Cryptochrome 2 variants engineered for different orders of cluster formation. This is exemplified by clustering eGFP-IKKα and eGFP-IKKβ, thereby achieving potent and reversible activation of NF-κB signaling. It is demonstrated that this approach can activate downstream signaling via the endogenous NF-κB pathway and is thereby capable of activating both an NF-κB-responsive reporter construct as well as endogenous NF-κB-responsive target genes as analyzed by RNA sequencing. The generic design of this system is likely transferable to other signaling pathways to analyze the kinetics of signal activation and propagation.
Ostmann, Katharina | Kraegeloh, Annette | Weber, Wilfried
DOI:
Formaldehyde is the smallest existing aldehyde, a highly reactive color less gas at room temperature and ubiquitously present in our atmosphere. Because of its reactivity leading to the crosslinking of macromolecules like proteins, it is widely used in industrial applications, but also in cell biology in order to preserve cells and tissues for further analysis. In this work, we show that formaldehyde releasing solutions commonly used for fixation of cells, can diffuse via the gas phase to the neighboring well and influence signaling processes in the therein cultured cells. To analyze this effect, we utilized a stable reporter cell line for YAP signaling or a gene expression-based reporter for activation of the NF-κB pathway. We could show that next to formaldehyde, also glutaraldehyde and acetaldehyde were able to activate those signaling pathways. Additionally, especially the stable reporter cell line based on YAP signaling can also be used as sensor for bioavailable formaldehyde, being highly sensitive, easy to use, and reversible. The observed impact of formaldehyde on cellular signaling underscores the need for careful planning of experimental protocols and emphasizes the importance of implementing proper controls when utilizing this reagent in cellular signaling analyses.
Schmitt, Franz-Josef | Shah Mehmood, Amna | Tüting, Christian | Phan, Hoang Trong | Reisdorf, Judith | Rieder, Fabian | Golmohamadi, Farzin Ghane | Verma, Rajni | Kastritis, Panagiotis L. | Laufer, Jan
DOI:
The pH dependence of the absorption and (time-resolved) fluorescence of two red-shifted fluorescent proteins, mCardinal and mNeptune, was investigated. Decay-associated spectra were measured following fluorescence excitation at 470 nm in PBS buffer with a pH that ranged from 5.5 to 8.0. The fluorescence of both proteins shows two different decay components. mCardinal exhibits an increase in the long-lived fluorescence component with acidification from 1.34 ns at pH 8.0 to 1.62 ns at pH 5.5. An additional fast decay component with 0.64 ns at pH 8.0 up to 1.1 ns at pH 5.5 was found to be blue-shifted compared to the long-lived component. The fluorescence lifetime of mNeptune is insensitive to pH. DAS of mCardinal were simulated assuming a coupled two-level system to describe the 1S state of the chromophore within two different conformations of the protein. MD simulations were conducted to correlate the experimentally observed pH-induced change in the lifetime in mCardinal with its molecular properties. While the chromophores of both protein variants are stabilized by the same number of hydrogen bonds, it was found that the chromophore in mCardinal exhibits more water contacts compared to mNeptune. In mCardinal, interaction between the chromophore and Glu-145 is reduced as compared to mNeptune, but interaction with Thr-147 which is Ser-147 in mNeptune is stronger in mCardinal. Therefore, the dynamics of the excited-state proton transfer (ESPT) might be different in mCardinal and mNeptune. The pH dependency of ESPT is suggested as a key mechanism for pH sensitivity.
Armbruster, Anja | Ehret, Anna K. | Russ, Marissa | Idstein, Vincent | Klenzendorf, Melissa | Gaspar, Denise | Juraske, Claudia | Yousefi, O. Sascha | Schamel, Wolfang W. | Weber, Wilfried | Hörner, Maximilian
DOI:
Optogenetics is a versatile and powerful tool for the control and analysis of cellular signaling processes. The activation of cellular receptors by light using optogenetic switches usually requires genetic manipulation of cells. However, this considerably limits the application in primary, nonengineered cells, which is crucial for the study of physiological signaling processes and for controlling cell fate and function for therapeutic purposes. To overcome this limitation, we developed a system for the light-dependent extracellular activation of cell surface receptors of nonengineered cells termed OptoREACT (Optogenetic Receptor Activation) based on the light-dependent protein interaction of A. thaliana phytochrome B (PhyB) with PIF6. In the OptoREACT system, a PIF6-coupled antibody fragment binds the T cell receptor (TCR) of Jurkat or primary human T cells, which upon illumination is bound by clustered phytochrome B to induce receptor oligomerization and activation. For clustering of PhyB, we either used tetramerization by streptavidin or immobilized PhyB on the surface of cells to emulate the interaction of a T cell with an antigen-presenting cell. We anticipate that this extracellular optogenetic approach will be applicable for the light-controlled activation of further cell surface receptors in primary, nonengineered cells for versatile applications in fundamental and applied research.
Fischer, Alexandra A. M. | Robertson, Hanah B. | Kong, Deqiang | Grimm, Merlin M. | Grether, Jakob | Groth, Johanna | Baltes, Carsten | Fliegauf, Manfred | Lautenschlaeger, Franziska | Grimbacher, Bodo | Ye, Haifeng | Helms, Volkhard | Weber, Wilfried
DOI:
Phase separation of biomolecules into condensates is a key mechanism in the spatiotemporal organization of biochemical processes in cells. However, the impact of the material properties of biomolecular condensates on important processes, such as the control of gene expression, remains largely elusive. Here, the material properties of optogenetically induced transcription factor condensates are systematically tuned, and probed for their impact on the activation of target promoters. It is demonstrated that transcription factors in rather liquid condensates correlate with increased gene expression levels, whereas stiffer transcription factor condensates correlate with the opposite effect, reduced activation of gene expression. The broad nature of these findings is demonstrated in mammalian cells and mice, as well as by using different synthetic and natural transcription factors. These effects are observed for both transgenic and cell-endogenous promoters. The findings provide a novel materials-based layer in the control of gene expression, which opens novel opportunities in optogenetic engineering and synthetic biology.

