We engineer cells and materials that communicate and process information through synthetic biology
Our inspiration is the ability of organisms and the materials they are made of to adapt to dynamic environmental conditions. Plants adapt growth to light conditions; bacteria develop resistance against antibiotics or bones get stronger when exercised. The basis for this ability to adapt is a fascinating information processing machinery of the organisms: Environmental conditions are captured by molecular sensors, then the signals are processed and integrated with genetic programs to finally yield a targeted response.
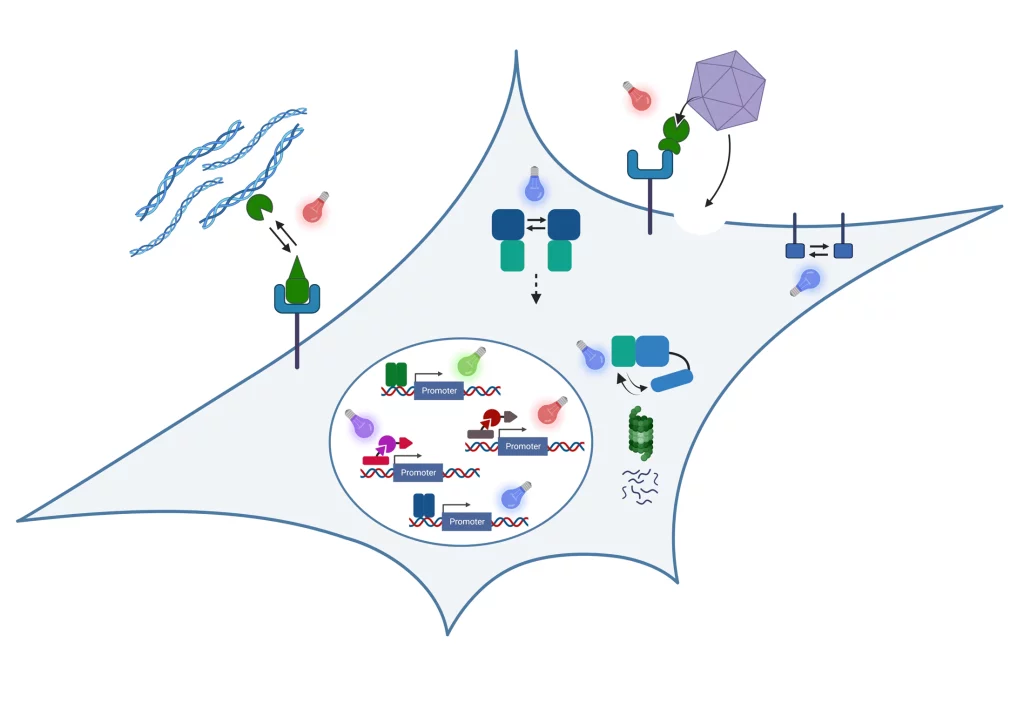
In our research, we engineer nature’s molecular sensing, processing, and actuation machinery in order to precisely control the function and properties of cells and materials. We apply these newly developed technologies in different fields of fundamental and applied research.

Team Members










Research
Stimulus-responsive and Information-processing (living) Materials

We develop and apply stimulus-responsive and information-processing biohybrid polymer materials. To this aim, we functionally couple synthetic biological molecular sensors and switches to polymer materials. By wiring these switches according to topologies inspired by electronic circuits, we engineer materials that perform fundamental computational operations. Examples of our work include:
- We engineered a hydrogel based on a bacteria-derived photoreceptor which allows the light-responsive, fully reversibly tuning of its mechanical properties. We applied this hydrogel as extracellular matrix to analyze the impact of dynamic mechanical environments on transcriptome-wide responses in mesenchymal stem cells or on the migration of T-lymphocytes.
See Hörner et al. Advanced Materials 2019 - We integrated synthetic biological switches with polymer materials into a circuit inspired by an electronic counter. The resulting material system was able to count the number of input light pulses and to release different output as a function of the number of light pulses detected. We applied this system to sequentially release different biocatalysts to drive a two-step biochemical reaction.
See Beyer et al., Advanced Materials 2018 - We developed PenTag, a protein tag for the spontaneous, covalent coupling of proteins to ampicillin-functionalized molecules such as dyes, polymers, or solid supports. Based on this strategy, we engineered and assembled material modules to function as encoder for processing different combinations of biochemical input stimuli.
See Mohsenin et al., Advanced Functional Materials 2024 - By engineering modular protease-based switches that can either be activated or repressed, we develop information-processing biohybrid circuits that process binary biomolecular information according to a circuit inspired by electronic decoders. Such circuits can be applied to process and interpret biochemical sensor information for advanced diagnostic applications.
See Mohsenin et al., Advanced Materials 2024
Molecular optogenetics to control cell fate and function
We develop and apply molecular optogenetic tools to control cell fate and function with unprecedented spatial and temporal precision in a dose-dependent and highly specific manner. To this aim, we engineer plant- and bacteria-derived photoreceptors and functionally couple them to proteins involved in cell signaling and gene expression. Examples of our work include:
- Light-inducible formation of liquid or gel-like transcription factor condensates in mammalian cells and mice. We demonstrate that liquid “transcription factor droplets” show a several-fold higher activity in inducing transgene expression compared to native transcription factors. Further, gel-like transcription factor condensates were shown to correlate with decreased transcriptional activation thus providing a materials-based layer of controlling gene expression.
See Schneider et al., Science Advances 2021 and Fischer et al., Small 2024 - Light-guided adeno-associated viral (AAV) vectors. We engineered a light-responsive tropism into AAVs which allows us to selectively transfer genetic information into single cells or to transduce different cells within one culture with different transgenes.
See Hörner et al., Science Advances 2021
Our group is running www.optobase.org, the most comprehensive database on molecular optogenetics. Have a look and discover the amazing opportunities in controlling biology with light!
Biosensors
We integrate natural and engineered molecular sensors for drugs, metabolites or nucleic acids into suitable readout formats for the fast and sensitive quantification of such substances. Together with collaboration partners, we develop biosensor systems for different application fields:
Open Positions
We are always excited to meet curious and creative scientists passionate about synthetic biology, optogenetics, and engineered living materials. If you would like to shape the future of biobased and living materials with us, we warmly welcome your spontaneous application for a PhD thesis or Postdoc position!
Projects and Partners
We perform collaborative research in materials-oriented synthetic biology within interdisciplinary research consortia
STEADY
Within the ERC Advanced Grant STEADY, we develop concepts for dynamically controlling the properties of engineered living materials by advanced synthetic genetic circuits.
LoopOfFun
We coordinate the European Innovation Council (EIC)-funded consortium LoopOfFun in which we aim at developing a platform for the rapid development of industry-scale, one-step, simple casting-based manufacturing processes for fungal mycelia composites. We jointly work towards this goal with our consortium partners:
- Prof. Roman Jerala, National Institute of Chemistry, Ljubljana, Slovenia
- Dr. Achim Weber, Fraunhofer IGB, Stuttgart, Germany
- Prof. Arnold Driessen, University of Groningen, The Netherlands
- Carlotta Borgato and Jan Boelen, Atelier LUMA, Arles, France
DELIVER
In the project DELIVER funded by the Carl-Zeiss-Foundation, we collaborate towards the data-driven engineering of sustainable living materials. We combine synthetic biology with materials sciences and data-driven approaches to design bio-based composite materials with custom-tailored structural properties for construction applications. Within deliver, we collaborate with the following partners:
- Prof. Thomas Speck, University of Freiburg, Germany
- Dr. Clemens Kreutz, University Hospital Freiburg, Germany
BILLARD
We coordinate the BILLARD project funded by the Federal Ministry of Education and Research (BMBF) within the funding line “Biologization of Technology”, we collaborate with PD Dr. Felicitas Bucher from the Clinic of Ophtamology at the University Hospital Freiburg on the development of novel intraocular drug delivery devices.
CIBSS – Centre for Integrative Biological Signalling Studies
We are member of the Cluster of Excellence CIBSS in which we perform research on novel optogenetic technologies to control signaling reactions in mammalian cells. We mainly collaborate with Prof. Dr. Jens Timmer on the model-based design of synthetic biological switches and networks and with Prof. Dr. Wolfgang Schamel on controlling immunological processes such as T cell activation via optogenetics.
Publications
Kuharenko, Olga V. | Antanovich, Artsiom | Saha, Avijit | Ivanchanka, Aliaksei | Müller, Martin | Lenyak, Vladimir | Kraegeloh, Annette | Rossner, Christian
DOI:
We report a straightforward methodology to access structurally well-defined hybrid assemblies of plasmonic and excitonic nanoparticles (NPs). The developed strategy is based on the incorporation of quantum dots (QDs) coated with zinc-sulfide shells into poly(ethylene glycol) (PEG) brushes at gold NP surfaces, without the necessity of incorporating specialized functional groups to drive the supracolloidal assembly. Based on control experiments involving PEGs with distinct polymeric architecture and Fourier-transform infrared spectroscopy analysis, we attribute the structure formation to attractive interactions between the QD surface and the monomeric repeat unit of the PEG brushes. This combination leads to short interparticle spacings and plasmon/exciton interactions, resulting in photoluminescence (PL) quenching upon assembly. However, using block-copolymers comprising a NP-adjacent spacer block in addition to a NP-remote PEG block, the distance between gold NPs and QDs can be controlled, which in turn affects the PL properties. The versatility of the structure-formation approach is demonstrated by the possibility of applying it to two distinct core/shell QDs (InP/ZnSe/ZnS and CdSe/CdS/ZnS). This offers new perspectives in the quest for efficient nanomaterial fabrication procedures.
Armbruster, Anja | Hörner, Maximilian | Wagner, Hanna J. | Fink-Straube, Claudia | Weber, Wilfried
DOI:
Recombinant adeno-associated viral (rAAV) vectors are a leading platform for in vivo gene therapy, valued for their excellent safety, broad serotype diversity, and scalable production. Targeted delivery through capsid display of ligands holds great promise, yet current retargeting strategies often rely on extensive capsid re-engineering and restrict the use of ligands incompatible with intracellular expression systems. Here, we present a modular AAV retargeting platform that, for the first time, employs the SpyTag/SpyCatcher system via genetic integration into the AAV2 capsid. SpyTag is a small peptide that forms a covalent, irreversible bond with its protein partner, SpyCatcher, allowing site-specific ligand coupling under physiological conditions. Inserting SpyTag into surface-exposed capsid sites enabled postassembly functionalization of AAVs with SpyCatcher-fused targeting proteins. As proof of concept, we used SpyCatcher fusions with designed ankyrin repeat proteins (DARPins) specific for EGFR, EpCAM, and HER2. This conferred highly specific transduction of corresponding cancer cell lines with minimal off-target activity. Therapeutic potential was demonstrated by delivering a suicide gene, inducing selective cancer cell killing upon prodrug administration. This “one-fits-all” platform allows rapid and flexible retargeting without significantly altering the underlying vectors genome or production process. It supports the incorporation of large or complex ligands not amenable to genetic fusion and facilitates high-throughput preclinical evaluation strategies. By uniting capsid engineering with modular ligand display, our approach provides a scalable and versatile framework for precision gene delivery, broadening the applicability of rAAV in both therapeutic and discovery settings.
Pirritano, Marcello | Buescher, Johannes | Staubach, Pauline | Tacken, Thorsten | Yakovleva, Yulia | Sabura, Mark | Shehu, Kristela | Franzenburg, Sören | Schneider, Marc
DOI:
Discrimination of self from non-self RNA is a critical requirement for any cell to respond to infections and to maintain cellular integrity. We report novel functions for two RNA-dependent RNA polymerases (RDRs) in Paramecium. In RNAinterference (RNAi), RDRs are normally involved in the production of large amounts of secondary small interfering RNAs (siRNAs). To characterize the function of RDRs in context of exogenous RNA recognition, we developed a novel double-stranded RNA (dsRNA) application system using dextran nanoparticles to deliver heteroduplex dsRNA to cells as food particles, mimicking the natural phagosomal entry. Small RNA sequencing allows to dissect siRNAs produced from exogenous RNA or RDR transcripts. Contrary to expectations, our data show that Dicer is unable to directly cleave exogenous dsRNA while two RDRs, RDR1 and RDR2, are required for the initial steps of dsRNA-induced RNAi. Paradoxically, these two RDRs must replicate dsRNA before Dicer cleavage. This system works efficiently also with exogenous single-stranded RNA (ssRNA), although RDR2 is dispensable for ssRNA conversion. The function of RDRs is in contrast to that in animals, plants and fungi and extends the functional diversity of these polymerases as RDR-associated complexes appear to control the entry of food RNA into the RNAi machinery.
Mohsenin, Hasti | Schmachtenberg, Rosanne | Kemmer, Svenja | Wagner, Hanna J. | Johnston, Midori | Madlener, Sibylle | Dincer, Can | Timmer, Jens | Weber, Wilfried
DOI:
The functional integration of biological switches with synthetic building blocks enables the design of modular, stimulus-responsive biohybrid materials. By connecting the individual modules via diffusible signals, information-processing circuits can be designed. Such systems are, however, mostly limited to respond to either small molecules, proteins, or optical input thus limiting the sensing and application scope of the material circuits. Here, a highly modular biohybrid material is design based on CRISPR/Cas13a to translate arbitrary single-stranded RNAs into a biomolecular material response. This system exemplified by the development of a cascade of communicating materials that can detect the tumor biomarker microRNA miR19b in patient samples or sequences specific for SARS-CoV. Specificity of the system is further demonstrated by discriminating between input miRNA sequences with single-nucleotide differences. To quantitatively understand information processing in the materials cascade, a mathematical model is developed. The model is used to guide systems design for enhancing signal amplification functionality of the overall materials system. The newly designed modular materials can be used to interface desired RNA input with stimulus-responsive and information-processing materials for building point-of-care suitable sensors as well as multi-input diagnostic systems with integrated data processing and interpretation.
Shehu, Kristela | Schneider, Marc | Kraegeloh, Annette
DOI:
Antibiotic resistance in chronic lung infections caused by Pseudomonas aeruginosa requires alternative approaches to improve antibiotic efficacy. One promising approach is the use of adjuvant compounds that complement antibiotic therapy. This study explores the potential of menadione as an adjuvant to azithromycin against planktonic cells and biofilms of P. aeruginosa, focusing on its mechanisms of action and cytotoxicity in pulmonary cell models. Methods: The effect of menadione in improving the antibacterial and antibiofilm potency of azithromycin was tested against P. aeruginosa. Mechanistic studies in P. aeruginosa and AZMr-E. coli DH5α were performed to probe reactive oxygen species (ROS) production and bacterial membrane disruption. Cytotoxicity of antibacterial concentrations of menadione was assessed by measuring ROS levels and membrane integrity in Calu-3 and A549 lung epithelial cells. Results: Adding 0.5 µg/mL menadione to azithromycin reduced the minimum inhibitory concentration (MIC) by four-fold and the minimum biofilm eradication concentration (MBEC) by two-fold against P. aeruginosa. Adjuvant mechanisms of menadione involved ROS production and disruption of bacterial membranes. Cytotoxicity tests revealed that antibacterial concentrations of menadione (≤64 µg/mL) did not affect ROS levels or membrane integrity in lung cell lines. Conclusions: Menadione enhanced the efficacy of azithromycin against P. aeruginosa while exhibiting a favorable safety profile in lung epithelial cells at antibacterial concentrations. These findings suggest that menadione is a promising antibiotic adjuvant. However, as relevant data on the toxicity of menadione is sparse, further toxicity studies are required to ensure its safe use in complementing antibiotic therapy.
Khazem, Ali | Schmachtenberg, Rosanne | Weiand, Anke | Sankaran, Shrikrishnan | Weber, Wilfried
DOI:
Living therapeutic and diagnostic materials based on engineered microorganisms are emerging as a novel approach with the perspective of providing patient-tailored, sustainable, and cost-effective healthcare solutions. In this review, we focus on recent advances in using genetically or chemically engineered microorganisms as living diagnostics, therapeutics, and as a means of prevention for various diseases. We also highlight the applications of living therapeutics for acute and chronic diseases, and the role of micro/macro-encapsulation of the engineered microorganisms. We further showcase the current success of engineered living therapeutics in clinical trials and discuss challenges and future trends in the field.
Jerez-Longres, Carolina | Weber, Wilfried
DOI:
Living natural materials have remarkable sensing abilities that translate external cues into functional changes of the material. The reconstruction of such sensing materials in bottom-up synthetic biology provides the opportunity to develop synthetic materials with life-like sensing and adaptation ability. Key to such functions are material modules that translate specific input signals into a biomolecular response. Here, we engineer a synthetic organelle based on liquid–liquid phase separation that translates a metabolic signal into the regulation of gene transcription. To this aim, we engineer the pyruvate-dependent repressor PdhR to undergo liquid–liquid phase separation in vitro by fusion to intrinsically disordered regions. We demonstrate that the resulting coacervates bind DNA harboring PdhR-responsive operator sites in a pyruvate dose-dependent and reversible manner. We observed that the activity of transcription units on the DNA was strongly attenuated following recruitment to the coacervates. However, the addition of pyruvate resulted in a reversible and dose-dependent reconstitution of transcriptional activity. The coacervate-based synthetic organelles linking metabolic cues to transcriptional signals represent a materials approach to confer stimulus responsiveness to minimal bottom-up synthetic biological systems and open opportunities in materials for sensor applications.
Schneider, Sophie H.E. | Lehnert, Kathrin | Thome, Marie A. | Kraegeloh, Annette | Lienkamp, Karen
DOI:
Materials that can be switched between a polycationic/antimicrobial and a polyzwitterionic/protein-repellent state have important applications, e.g., as biofilm-reducing coatings in medical devices. However, the lack of stability under storage and application conditions so far restricts the lifetime and efficiency of such materials. In this work, a polynorbornene-based polycarboxybetaine with an optimized molecular structure for improved hydrolytic stability is presented. The polymer is fully characterized on the molecular level. Surface-attached polymer networks are obtained by spin-coating and UV cross-linking. These coatings are highly uniform and demonstrate charge-switching in zeta-potential studies. Storage stability in the dry state, as well as in aqueous systems at pH 4.5 and 7.4 for 28 days, is demonstrated. At pH 8, hydrolytic degradation is observed. Overall, the materials are substantially more stable than the corresponding ester-based systems.
Mohamed, Islam | Burckhardt, Kristin | Lohse, Stefan
DOI:
Neutrophils are innate immune cells that perpetually patrol the circulation and tissues. They sense and migrate toward invading microbes to initiate and orchestrate a robust immune response. Their highly reactive nature, driven by multiple and redundant receptor families recognizing bacterial components, makes them particularly sensitive to contaminants or nonsterile implants. This often leads to a neutrophil-driven foreign body reaction that shields the implant and triggers inflammation, collateral tissue damage, or even sepsis. This presents a significant challenge for living therapeutic materials, an innovative biomedical approach using genetically engineered bacteria encapsulated in natural or synthetic polymers. Since bacterial turnover inevitably releases pathogen-associated molecular patterns that activate neutrophils to mitigate or prevent a potent neutrophil response, living therapeutic material design strategies are required to protect the living therapeutic material from damage while maintaining its functionality. This review focuses on current strategies involving bacterial genetic engineering, immune-shielding materials and factors, and modified hydrogel-based systems to minimize immune recognition. Engineering the bacterial chassis to produce immune tolerance–inducing metabolites from commensals, modified pathogen-associated molecular patterns, and pathogen-associated molecular pattern–cleaving autolysins may enhance biocompatibility. A crucial aspect for clinical translation is robust biocontainment to prevent bacterial escape, ensuring living therapeutic material remains a safe and effective therapeutic platform. While the potential of the living therapeutic material concept lies in the development of tailored medicine specifically designed for a specific disease and enabling local, cost-effective, site- and stimulus-responsive treatment, balancing the neutrophil immune response remains an important milestone on the path to living therapeutic material for future biomedical applications.
Heilmann, Heiko | Busch, Lukas | Buchmann, Celine | Mohamed, Islam | Theiß, Adrian | Junkger, Sabryna | Lohse, Stefan | Bufe, Bernd
DOI:
Formyl peptide receptors (FPRs) are pattern recognition receptors well-known for bacterial pathogen sensing. We here identified activator and inhibitor motifs for FPRs that are present on surface proteins of various viral pathogens. Peptides containing these motifs interact with all FPR family members and modulate various important immune functions in innate immune cells. Viral breakdown products comprising these motifs were found in patients with COVID-19. In the spike protein, many activators are found in highly mutagenic regions, whereas the inhibitor motif is located in a conserved domain that also exists in further unrelated viruses. The physiochemical properties of FPR1 activators correlate with the occurrence of protein aggregation hotspots. Such hotspots are present on various surface proteins of unrelated viruses that can also activate FPRs. This points toward a general contribution of FPRs in modulating antiviral immune responses during many distinct viral infections.


