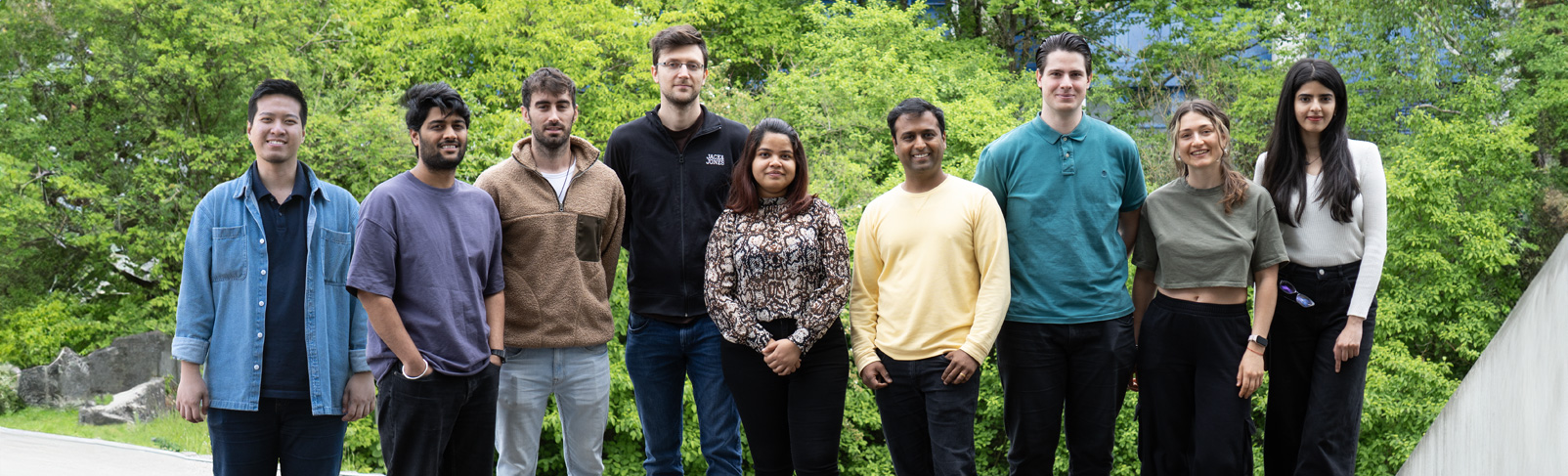
The Research Group Bioprogrammable Materials explores a young multidisciplinary field combining synthetic biology and biomaterials. It focuses on the development of materials with genetically programmed functionalities capable of biosensing, stimuli-responsive long-term drug release, and manipulation of cell behavior. Synthetic biology tools are used to program proteins and microbes to perform smart and beneficial functions. These engineered biological entities are then incorporated in appropriately developed polymeric matrices, resulting in composite materials with highly versatile functionalities, a wide range of tunability, and in situ controllability.
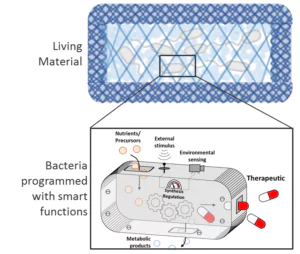
Figure: Bacterias are programmed with smart functions for applications e.g. in biosensing and drug delivery.

Staff








Research
In the Bioprogrammable Materials group, we combine genetically-programmed living organisms like bacteria with polymeric matrices like hydrogels to generate smart composite materials called engineered living materials (ELMs). Our ELMs are designed for a range of biomedical applications, such as biosensing and drug-delivery through innovations in both the living and non-living components:
Stimuli-responsive drug-secretion in bacteria
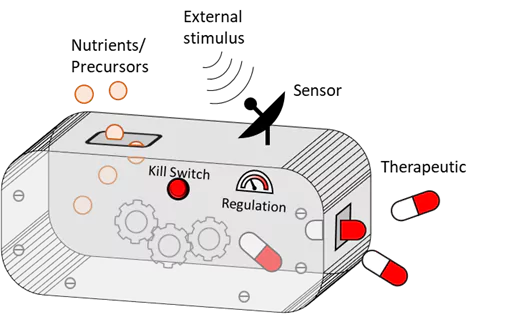
Bacteria are an integral part of the body’s microbiome, with several native and probiotic species imparting natural health benefits to humans. Bacteria are also used extensively in the pharmaceutical industry as biofactories to produce a variety of drugs. Our research merges these two features by engineering medically relevant bacteria like E. coli, Lactobacilli and Corynebacterium to produce and secrete therapeutic molecules directly in the body where they are needed. Since the bacteria naturally thrive within the body, long-term drug release can be sustained. We engineer the bacteria to produce and release anti-microbial, anti-inflammatory and regenerative drugs in the form of peptides, proteins and enzymatically synthesized biomolecules to treat chronic diseases.
Due to variabilities in patient profiles and disease progressions, it is highly desirable to personalize and customize the activity of these therapeutic bacteria in order to ensure their effectiveness. For this, we are developing genetic modules which allow external stimuli to “switch on” drug release. Stimuli like light, heat and small molecules allow remote-control over the bacteria, and stimuli like disease biomarkers will enable autoregulation of bacterial activity in response to disease progression. Stimuli-responsive genetic modules are also being developed as an additional layer of biosafety, to ensure that bacteria do not survive outside their intended implant environment. While many of the genetic modules we use were developed for E. coli, we are also endowing probiotic bacteria like lactobacilli and Corynebacterium with similar functions – a challenge which promises significant rewards by expanding the range of ELM-ready organisms and functions.
Recent publications:
- Dhakane, P.; Tadimarri, V. S.; Sankaran, S. Light-Regulated pro-Angiogenic Engineered Living Materials. bioRxiv – https://doi.org/10.1101/2022.10.28.514190.
- Dey, S.; Asensio, M. B.; Kuttae, S. B.; Sankaran, S. Novel Genetic Modules Encoding High-Level Antibiotic-Free Protein Expression in Probiotic Lactobacilli. bioRxiv – https://doi.org/10.1101/2022.08.04.502766.
Hydrogels for bacterial encapsulation and tools to understand bacterial behavior in confinement
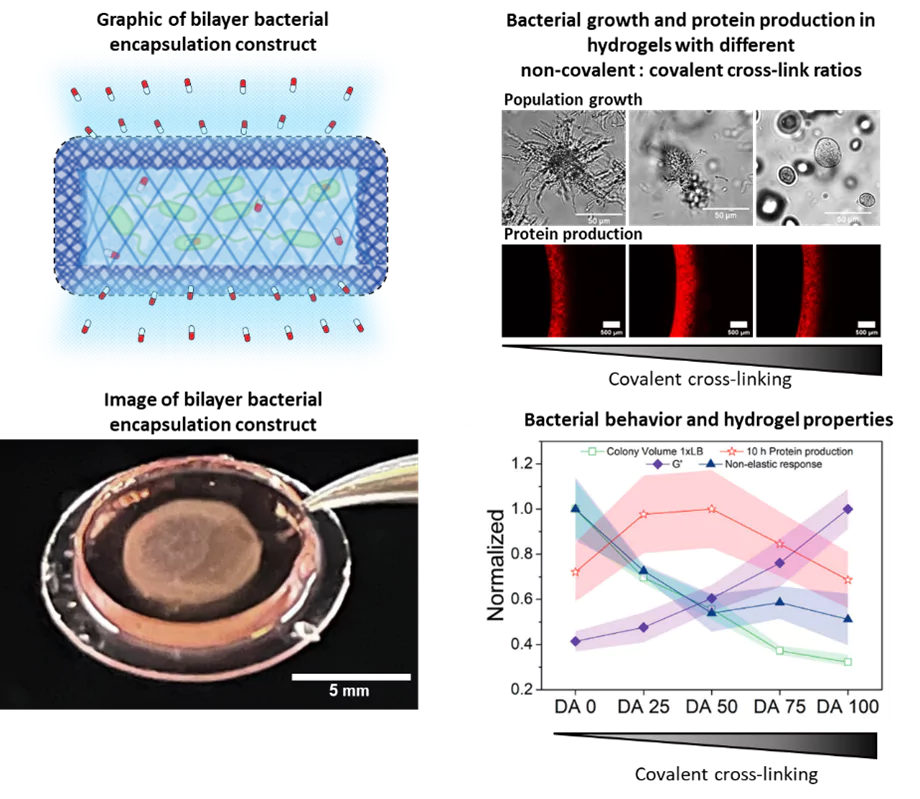
For the therapeutic bacteria to treat a disease, they need to colonize the disease site and establish a population big enough to provide effective drug doses. In collaboration with the Dynamic Biomaterials group, we are developing hydrogel matrices that provide conducive environments for the bacteria to grow and function. These hydrogels are designed to mechanically control the size of the contained bacterial population while maximizing their functionality. The hydrogel network allows diffusion of nutrients, metabolites and drugs in and out of the matrix, while preventing bacterial escape and providing protection from immune cells. The material component therefore provides an added level of biosafety for the use of genetically modified bacteria within the body.
As part of this research, we have identified that certain mechanical properties of the hydrogels influence growth and metabolism of the bacteria within, similar to how it occurs in natural biofilms. This, in turn, affects the performance of the bacteria in terms of stimuli-responsiveness and drug production. Thus, we are studying bacterial behavior confined in hydrogels, whole mechanical properties can be tuned. Thereby, we are unravelling the interplay between collective bacterial behavior and mechanical forces surrounding them. Apart from this enabling us to optimize the performance of our engineered living materials, it also serves as an artificial biofilm platform to gain fundamental insights into potential behavior of bacteria within different domains of natural biofilms. We use microscopy, biochemical and genetic assays to determine growth and metabolic behavior of the bacteria over time.
Recent publications:
Bhusari, S.; Sankaran, S.; del Campo, A. Regulating Bacterial Behavior within Hydrogels of Tunable Viscoelasticity. Advanced Science 2022, 9 (17), 2106026. https://doi.org/10.1002/advs.202106026.
Bhusari, S.; Kim, J.; Polizzi, K.; Sankaran, S.; Campo, A. del. Encapsulation of Bacteria in Bilayer Pluronic Thin Film Hydrogels: A Safe Format for Engineered Living Materials. bioRxiv – https://doi.org/10.1101/2022.09.29.510162.
Collaborations:
These research endeavors are also part of multiple collaborations within two consortia – (i) Leibniz Science Campus on Living Therapeutic Materials (LSC LifeMat) and (ii) Collective Research Center on Physical modeling of non-equilibrium processes in biological systems (CRC 1027).
Finanzierung:


Publications
Blanch-Asensio, Marc | Tadimarri, Varun S. | Wilk, Alina | Sankaran, Shrikrishnan
DOI:
Background: The Lactobacillaceae family comprises many species of great importance for the food and healthcare industries, with numerous strains identified as beneficial for humans and used as probiotics. Hence, there is a growing interest in engineering these probiotic bacteria as live biotherapeutics for animals and humans. However, the genetic parts needed to regulate gene expression in these bacteria remain limited compared to model bacteria like E. coli or B. subtilis. To address this deficit, in this study, we selected and tested several bacteriophage-derived genetic parts with the potential to regulate transcription in lactobacilli.
Results: We screened genetic parts from 6 different lactobacilli-infecting phages and identified one promoter/repressor system with unprecedented functionality in Lactiplantibacillus plantarum WCFS1. The phage-derived promoter was found to achieve expression levels nearly 9-fold higher than the previously reported strongest promoter in this strain and the repressor was able to almost completely repress this expression by reducing it nearly 500-fold.
Conclusions: The new parts and insights gained from their engineering will enhance the genetic programmability of lactobacilli for healthcare and industrial applications.

Dey, Sourik | Blanch-Asensio, Marc | Kuttae, Sanjana Balaji | Sankaran, Shrikrishnan
DOI:
Lactobacilli are ubiquitous in nature, often beneficially associated with animals as commensals and probiotics, and are extensively used in food fermentation. Due to this close-knit association, there is considerable interest to engineer them for healthcare applications in both humans and animals, for which high-performance and versatile genetic parts are greatly desired. For the first time, we describe two genetic modules in Lactiplantibacillus plantarum that achieve high-level gene expression using plasmids that can be retained without antibiotics, bacteriocins or genomic manipulations. These include (i) a promoter, PtlpA, from a phylogenetically distant bacterium, Salmonella typhimurium, which drives up to 5-fold higher level of gene expression compared to previously reported promoters and (ii) multiple toxin-antitoxin systems as a self-contained and easy-to-implement plasmid retention strategy that facilitates the engineering of tuneable transient genetically modified organisms. These modules and the fundamental factors underlying their functionality that are described in this work will greatly contribute to expanding the genetic programmability of lactobacilli for healthcare applications.
Riedel, Florian | Puertas Bartolomé, María | Teruel Enrico, Lara Luana | Fink-Straube, Claudia | Nguyen Dong, Cao | Gherlone, Fabio | Huang, Ying | Valiante, Vito | del Campo, Aránzazu | Sankaran, Shrikrishnan
DOI:
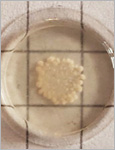
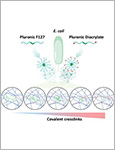
Microbial biofactories allow the upscaled production of high-value compounds in biotechnological processes. This is particularly advantageous for compounds like flavonoids that promote better health through their antioxidant, antibacterial, anticancer and other beneficial effects but are only produced in small quantities in their natural plant-based hosts. Bacteria like E. coli have been genetically modified with enzyme cascades to produce flavonoids like naringenin and pinocembrin from coumaric or cinnamic acid. Despite advancements in yield optimization, the production of these compounds still involves high costs associated with their biosynthesis, purification, storage and transport. An alternative production strategy could involve the direct delivery of the microbial biofactories to the body. In such a strategy, ensuring biocontainment of the engineered microbes in the body and controlling production rates are major challenges. In this study, these two aspects are addressed by developing engineered living materials (ELMs) consisting of probiotic microbial biofactories encapsulated in biocompatible hydrogels. Engineered probiotic E. coli Nissle 1917 able to efficiently convert cinnamic acid into pinocembrin were encapsulated in poly(vinyl alcohol)-based hydrogels. The biofactories are contained in the hydrogels for a month and remain metabolically active during this time. Control over production levels is achieved by the containment inside the material, which regulates bacteria growth, and by the amount of cinnamic acid in the medium.

Bhusari, Shardul | Sankaran, Shrikrishnan | del Campo, Aránzazu
DOI:
Engineered living materials (ELMs) are a new class of materials in which living organism incorporated into diffusive matrices uptake a fundamental role in material's composition and function. Understanding how the spatial confinement in 3D can regulate the behavior of the embedded cells is crucial to design and predict ELM's function, minimize their environmental impact and facilitate their translation into applied materials. This study investigates the growth and metabolic activity of bacteria within an associative hydrogel network (Pluronic-based) with mechanical properties that can be tuned by introducing a variable degree of acrylate crosslinks. Individual bacteria distributed in the hydrogel matrix at low density form functional colonies whose size is controlled by the extent of permanent crosslinks. With increasing stiffness and elastic response to deformation of the matrix, a decrease in colony volumes and an increase in their sphericity are observed. Protein production follows a different pattern with higher production yields occurring in networks with intermediate permanent crosslinking degrees. These results demonstrate that matrix design can be used to control and regulate the composition and function of ELMs containing microorganisms. Interestingly, design parameters for matrices to regulate bacteria behavior show similarities to those elucidated for 3D culture of mammalian cells.
Rodrigo-Navarro, Aleixandre | Sankaran, Shrikrishnan | Dalby, Matthew J. | del Campo, Aránzazu | Salmeron-Sanchez, Manuel
DOI:
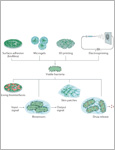
Biomaterials have evolved from inert materials that lack interaction with the body to biologically active, instructive materials that host and provide signals to surrounding cells and tissues. Engineered living materials contain living cells (responsive function) and polymeric matrices (scaffolding function) and, thus, can be designed as active and response biomaterials. In this Review, we discuss engineered living materials that incorporate microorganisms as the living, bioactive component. Microorganisms can provide complex responses to environmental stimuli, and they can be genetically engineered to allow user control over responses and integration of numerous inputs. The engineered microorganisms can either generate their own matrix, such as in biofilms, or they can be incorporated in matrices using various technologies, such as coating, 3D printing, spinning and microencapsulation. We highlight biomedical applications of such engineered living materials, including biosensing, wound healing, stem-cell-based tissue engineering and drug delivery, and provide an outlook to the challenges and future applications of engineered living materials.

