Curriculum Vitae
Education
2017 – 2020
PhD in Biology (summa cum laude), Heidelberg University
Max Planck Institute for Medical Research (Prof. Joachim P. Spatz)
2015 – 2017
M.Sc. Molecular Biotechnology, Heidelberg University
Max Planck Institute for Intelligent Systems, Stuttgart, Germany
2011 – 2015
B.Sc. Molecular Biotechnology, Heidelberg University
Dept. for Biophysical Chemistry, Heidelberg University, Germany
Academic Positions
Since 2022
Research Group Leader
Leibniz Institute for New Materials, Saarbrücken Germany
Since 2022
Adjunct Research Group Leader
Max Planck Bristol Center for Minimal Biology, Bristol, UK
2021 – 2022
Marie Skłodowska Curie Individual Postdoctoral Research Fellow
Kennedy Institute of Rheumatology, University of Oxford, UK
2020 – 2021
Postdoctoral Scientist
Max Planck Institute for Medical Research, Heidelberg, Germany
Awards, Fellowships and Recognitions
- Alois Lauer Research Prize for Medicine, Alois Lauer Foundation
- Member of Junge Akademie Mainz, Academy of Sciences and Literature Mainz
- Emmy Noether Research Group Leader, German Science Foundation
- Daimler and Benz Foundation, Postdoctoral Fellow
- Otto Hahn Medal, Max Planck Society
- Young Investigator Award, International Society of Extracellular Vesicles
- Karl-von-Frisch Award, German Life Science Organization (vBio)
- Individual Fellowship, Marie Skłodowska Curie Program, European Commission
- Walter Benjamin Postdoctoral Fellowship, German Science Foundation
- Add-on Postdoctoral Fellowship, Joachim Herz Foundation
- PhD-Fellowship, Max Planck School Matter to Life
- PhD-Fellowship, Hartmut Hoffmann-Berling Graduate School
- Exchange Fellowship, Elisabeth Meurer Foundation
- Study Fellowship, German National Merit Fundation (Studienstiftung des Deutschen Volkes)
Publications

Burgstaller, Anna | Piernitzki, Nils | Küchler, Nadja | Koch, Marcus | Kister, Thomas | Eichler, Hermann | Kraus, Tobias | Schwarz, Eva C. | Dustin, Michael | Lautenschlaeger, Franziska | Staufer, Oskar
DOI:
The expansion of T cells ex vivo is crucial for effective immunotherapy but currently limited by a lack of expansion approaches that closely mimic in vivo T cell activation. Taking inspiration from bottom-up synthetic biology, a new synthetic cell technology is introduced based on dispersed liquid-liquid phase-separated droplet-supported lipid bilayers (dsLBs) with tunable biochemical and biophysical characteristics, as artificial antigen presenting cells (aAPCs) for ex vivo T cell expansion. These findings obtained with the dsLB technology reveal three key insights: first, introducing laterally mobile stimulatory ligands on soft aAPCs promotes expansion of IL-4/IL-10 secreting regulatory CD8+ T cells, with a PD-1 negative phenotype, less prone to immune suppression. Second, it is demonstrated that lateral ligand mobility can mask differential T cell activation observed on substrates of varying stiffness. Third, dsLBs are applied to reveal a mechanosensitive component in bispecific Her2/CD3 T cell engager-mediated T cell activation. Based on these three insights, lateral ligand mobility, alongside receptor- and mechanosignaling, is proposed to be considered as a third crucial dimension for the design of ex vivo T cell expansion technologies.
DOI:
Synthetic cells can advance immunotherapy, offering innovative approaches to understanding and enhancing immune responses. This review article delves into the advancements and potential of synthetic cell technologies in immunology, emphasizing their role in understanding and manipulating immune functions. Recent progress in understanding vertebrate immune systems and the challenges posed by diseases highlight the need for innovative research methods, complementing the analysis of multidimensional datasets and genetic engineering. Synthetic immune cell engineering aims to simplify the complexity of immunological systems by reconstructing them in a controlled setting. This approach, alongside high-throughput strategies, facilitates systematic investigations into immunity and the development of novel treatments. The article reviews synthetic cell technologies, focusing on their alignment with the three laws of immunity: universality, tolerance, and appropriateness. It explores the integration of synthetic cell modules to mimic processes such as controlled T-cell activation, bacteria engulfment and elimination, or cellular maturation into desirable phenotypes. Together, such advancements expand the toolbox for understanding and manipulating immune functions. Synthetic cell technologies stand at the innovation crossroads in immunology, promising to illuminate fundamental immune system principles and open new avenues for research and therapy.
Buzas, Dora | Bunzel, Adrian | Staufer, Oskar | Milodowski, Emily J. | Edmunds, Grace L. | Bufton, Joshua | Vidana Mateo, Beatrice V. | Yadav, Sathish K.N. | Gupta, Kapil | Fletcher, Charlotte | Williamson, Maia K. | Harrison, Alexandra | Borucu, Ufuk | Capin, Julien | Francis, Ore | Balchin, Georgia | Hall, Sophie | Vega, Mirella V. | Durbesson, Fabien | Lingappa, Srikanth | Vincentelli, Renaud | Roe, Joe | Wooldridge, Linda | Burt, Rachel | Anderson, Ross J. L. | Mulholland, Adrian | Bristol UNCOVER Group | Hare, Jonathan | Bailey, Mick | Davidson, Andrew D. | Finn, Adam | Morgan, David | Mann, Jamie | Spatz, Joachim | Garzoni, Frederic
DOI:
Background: Due to COVID-19, pandemic preparedness emerges as a key imperative, necessitating new approaches to accelerate development of reagents against infectious pathogens. Methods: Here, we developed an integrated approach combining synthetic, computational and structural methods with in vitro antibody selection and in vivo immunization to design, produce and validate nature-inspired nanoparticle-based reagents against severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2).
Results: Our approach resulted in two innovations: (i) a thermostable nasal vaccine called ADDoCoV, displaying multiple copies of a SARS-CoV-2 receptor binding motif derived epitope and (ii) a multivalent nanoparticle superbinder, called Gigabody, against SARS-CoV-2 including immune-evasive variants of concern (VOCs). In vitro generated neutralizing nanobodies and electron cryo-microscopy established authenticity and accessibility of epitopes displayed by ADDoCoV. Gigabody comprising multimerized nanobodies prevented SARS-CoV-2 virion attachment with picomolar EC50. Vaccinating mice resulted in antibodies cross-reacting with VOCs including Delta and Omicron.
Conclusion: Our study elucidates Adenovirus-derived dodecamer (ADDomer)-based nanoparticles for use in active and passive immunization and provides a blueprint for crafting reagents to combat respiratory viral infections.
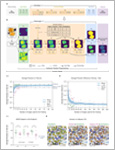

Cortacero, Kévin | McKenzie, Brienne | Müller, Sabina | Khazen, Roxana | Lafouresse, Fanny | Corsaut, Gaelle | Van Acker, Nathalie | Frenois, Francois-Xavier | Lamant, Laurence | Meyer, Nicolas | Vergier, Béatrice | Wilson, Dennis G. | Luga, Hervé | Staufer, Oskar | Dustin, Michael L. | Valitutti, Salvatore | Cussat-Blanc, Sylvain
DOI:
An unresolved issue in contemporary biomedicine is the overwhelming
number and diversity of complex images that require annotation, analysis and
interpretation. Recent advances in Deep Learning have revolutionized the field
of computer vision, creating algorithms that compete with human experts in
image segmentation tasks. However, these frameworks require large human-
annotated datasets for training and the resulting “black box” models are dif-
ficult to interpret. In this study, we introduce Kartezio, a modular Cartesian
Genetic Programming-based computational strategy that generates fully
transparent and easily interpretable image processing pipelines by iteratively
assembling and parameterizing computer vision functions. The pipelines thus
generated exhibit comparable precision to state-of-the-art Deep Learning
approaches on instance segmentation tasks, while requiring drastically smaller
training datasets. This Few-Shot Learning method confers tremendous flex-
ibility, speed, and functionality to this approach. We then deploy Kartezio to
solve a series of semantic and instance segmentation problems, and demon-
strate its utility across diverse images ranging from multiplexed tissue histo-
pathology images to high resolution microscopy images. While the flexibility,
robustness and practical utility of Kartezio make this fully explicable evolu-
tionary designer a potential game-changer in the field of biomedical image
processing, Kartezio remains complementary and potentially auxiliary to
mainstream Deep Learning approaches.

